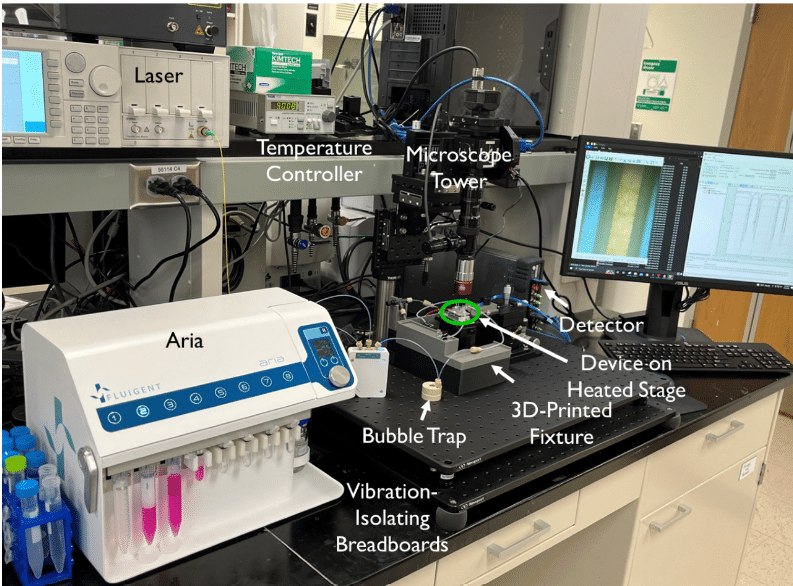
Microfluidic Drug Discovery
Microfluidic automated fluidic workflows as valuable tools for drug discovery
- Increased throughput and reproducibility
- Minimal reagent consumption
- Fast reaction times
- Automated fluidic workflow
Main applications in microfluidic drug discovery
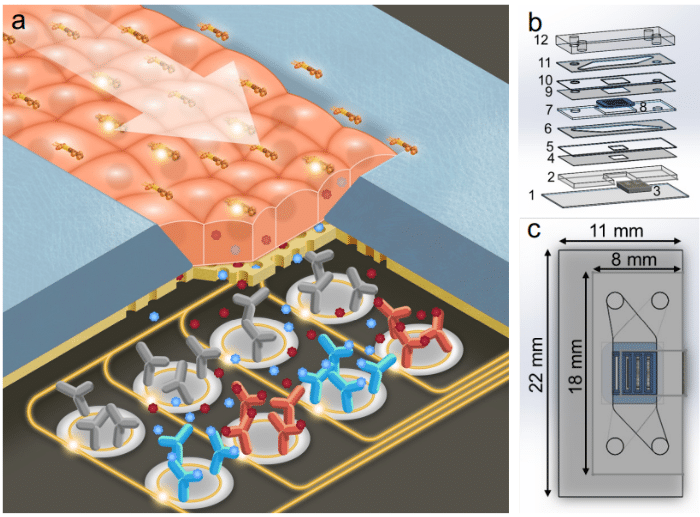
Target selection and validation (drug synthesis)
When developing a drug target, protein structural and affinity studies are needed for target selection and validation, and protein interactions within cells must be studied. Single-cell protein quantitation, protein analysis in nanoliter droplets, and high-sensitivity ligand-binding interactions can be performed using microfluidic devices.
Hit identification and optimization (drug screening)
In microfluidics for drug development, the potential pool size of drug candidates is estimated to be of the order of 1063. Microfluidic drug screening devices, including high-throughput microfluidic multiplexed systems that contain thousands of micrometer chambers, microwell arrays, or microvalves perform high-throughput screening studies with higher sensitivity and shorter reaction times while decreasing reagents volumes and costs.

Preclinical studies (drug evaluation)
In recent years, alternatives to animal experimentation have become a research hotspot. In microfluidic drug discovery, organs-on- chips and organoids are widely used as alternatives as they emulate the architectural and functional complexity of native organs. These systems also enable the exploration of facets of human disease and development that are not accurately recapitulated by animal models.
Resources
Flow Control Performance and Enhanced Automation for Microfluidic Drug Discovery Applications
- Drug synthesis and droplet microfluidics: When it comes to droplet or particle generation, having control of the fluid delivery system is important. During droplet or particle production, the flow rate of each phase must remain constant and stable to allow the production of monodisperse droplets.
- As experts in fluid control for droplet microfluidics, Fluigent provides droplet generation packages and platforms for researchers. These seamlessly generate micrometer droplets for a wide range of applications. Custom systems are also available to integrate technological and industrial devices.
- Multiplexing for drug screening: More than twenty reagents can be tested simultaneously when performing multiplexing applications on microfluidic devices. In microfluidic drug discovery, automated fluidic workflows are a prerequisite for drug screening. Fluigent developed the Aria to automate multiple fluid deliveries. Reproducibility is improved through complete automation of the fluidic protocol. Fluigent is also able to develop a custom system to fit user specifications and requirements.
- Optimized shear stress and recirculation for organ on a chip studies and cell culture: When performing drug evaluation on 2D/3D cell culture and organs-on-chips, cell culture should be optimized through constant perfusion to enable nutrients/oxygen renewal and by inducing adequate passive stimulation through shear flow as the effect is substantial on cell properties. Sterility on the fluidic path is also a prerequisite. Fluigent develops fluid recirculating systems that perform unidirectional fluid recirculation, ensuring improved and more stable flow rates compared to peristaltic pumps. The system can run for 10 days. Custom systems are also available for microfluidic drug discovery technology integration into industrial devices.
Fluigent Solutions as an Alternative to Syringe Pumps for Better Performance and Automation
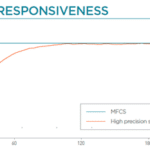
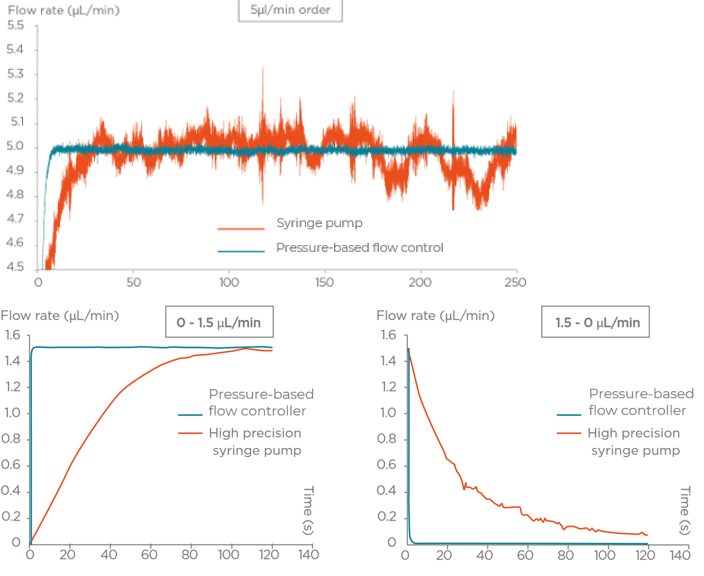
Flow rate stability and responsiveness is critical for the microfluidics for drug development applications. Syringe pumps are commonly used during this process. Depending on the model in use, syringe pumps show limited flow control, and the actual flow rate cannot be monitored. In addition, injection volumes are limited by the syringe, and automation can thus be difficult. An alternative to syringe pumps is pressure-based flow controllers. These show high-precision flow control, fast reaction time, and flow monitoring: parameters which are paramount for microfluidic drug discovery applications.
The benefits of choosing Fluigent for your flow control system:
- Best in class stability: < 0.5% thanks to our field-proven, patented FASTAB™ technology allowing optimal flow control with the robustness required in demanding industrial environments
- Workflow automation becomes straightforward with the Fluigent SDK and software included in the system
- Our engineering team members are experts in microfluidic design, mechanical and software integration, and biology applications
Related resources

5 reasons to choose OEM pressure controllers over OEM syringe pumps for microfluidic applications
Read moreOrgan on a chip Towards the next generation of cell culture platforms
Read more- Microfluidics Case Studies
OEM Case Study: Microfluidic Drug Screening
Read more - Expert Reviews: Basics of Microfluidics
Key reliability indicators for OEM components to ensure long-term performance of your flow control system
Read more - Expert Reviews: Basics of Microfluidics
Key considerations for fluidic system integration
Read more - Expert Reviews: Basics of Microfluidics
Prostate Organoid Culture in Microbeads
Read more - Expert Reviews: Basics of Microfluidics
Flow control for droplet generation using syringe pumps and pressure-based flow controllers
Read more - Expert Reviews: Basics of Microfluidics
Flow Control Technologies: Comparison between peristaltic, syringe and pressure pumps for microfluidic applications
Read more
Table of contents:
How Does Microfluidics Compare Classical Methods in Drug Delivery?
Traditional drug delivery methods, including oral administration, injections, and inhalation, have long depended on conventional techniques to fabricate drug carriers such as liposomes and polymeric nanoparticles.1,2 Although these systems aim to enhance drug solubility, stability, and targeting, their fabrication processes are often complex, difficult to scale, and limited in precision and reproducibility. Combined with challenges like poor biodistribution and biological barriers, these limitations reduce the effectiveness of many therapeutic agents.3
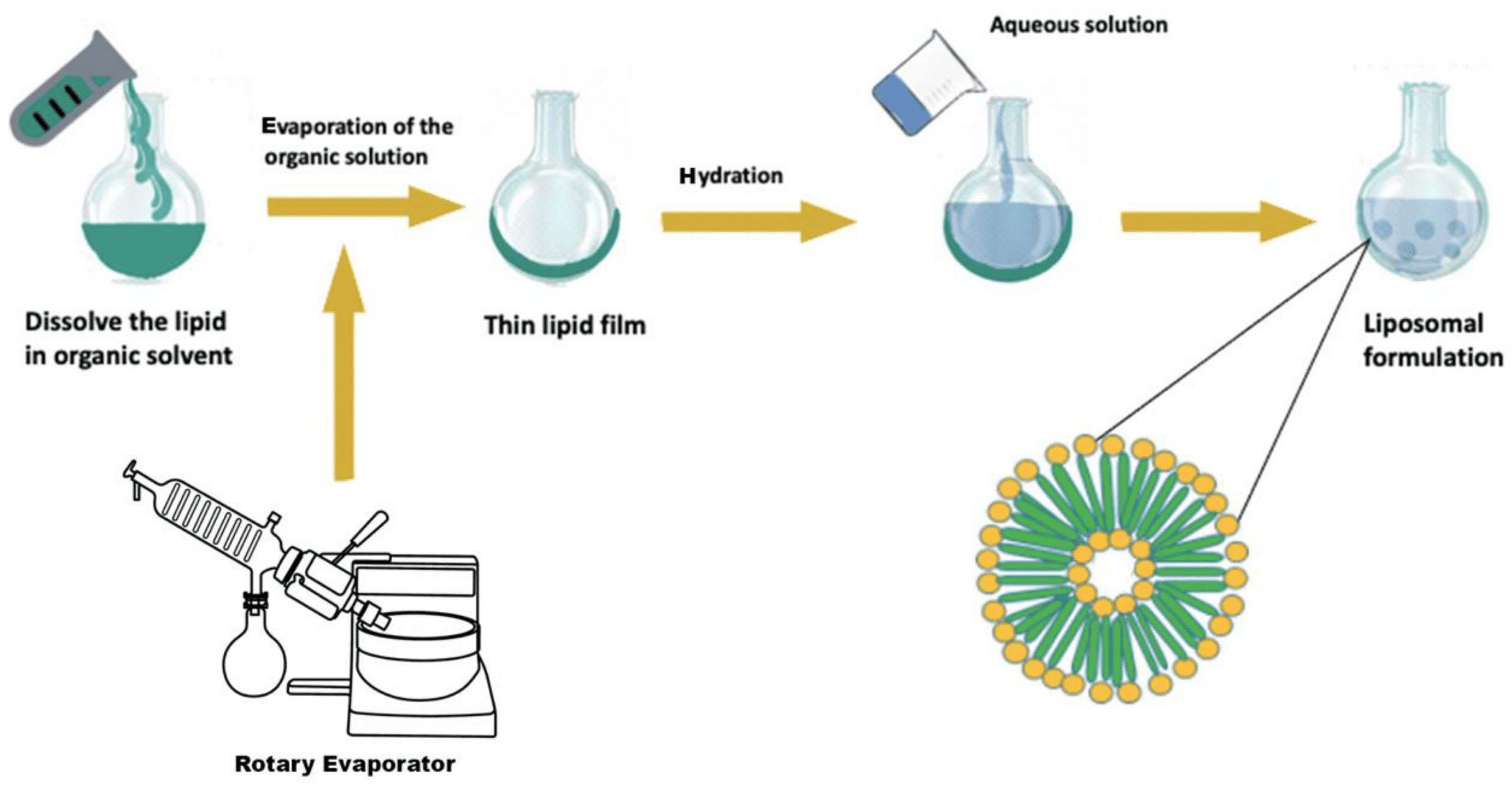
Microfluidic technology in healthcare offers a promising shift toward more efficient and controlled delivery systems. Known for its ability to manipulate fluids at the microscale, microfluidics enables the precise, scalable, and reproducible production of advanced drug carriers (Figure 1).4–6
This review explores recent developments in precision drug delivery using microfluidics, structured into three main areas:
(a) the fabrication of drug carriers, including lipid-based and polymeric nanoparticles, through microfluidic platforms;
(b) the integration of microfluidics into microneedle technologies for minimally invasive applications;
(c) the broader applications of microfluidics in nanomedicine and drug research, such as crystallization techniques and the development of in vitro platforms for studying delivery mechanisms.
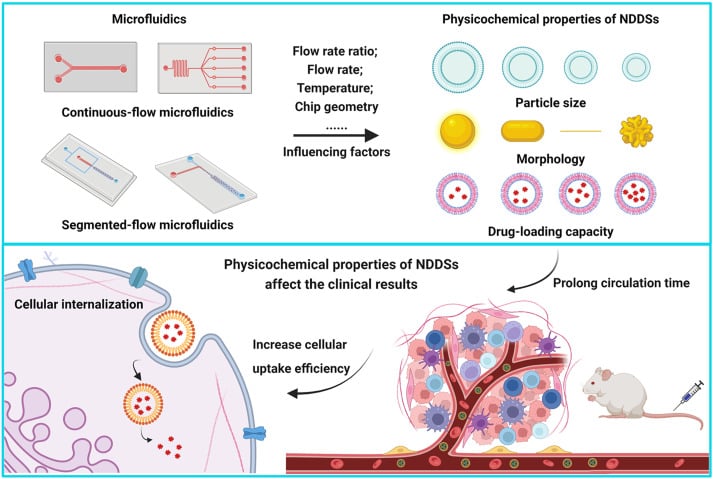
From Zhang, H. et al.; Acta Pharmaceutica Sinica B 2023, 13 (8), 3277–3299).
How Do Microfluidic Techniques Improve Drug Delivery Particle Synthesis?
1- Microfluidic Techniques for Lipid Nanoparticle Synthesis:
Lipid-based nanoparticles (LNPs) are a promising class of drug delivery systems due to their biocompatibility, capacity for encapsulating diverse therapeutic agents, and ability to facilitate controlled drug release. LNPs are particularly useful for delivering hydrophobic drugs, nucleic acids, and other sensitive molecules. Here are some microfluidic techniques used in LNP synthesis for drug delivery:7
- Microfluidic Hydrodynamic Focusing (MHF):
This technique uses a central stream of lipids dissolved in solvent, bordered by aqueous buffer streams. As the lipid solution is focused into a thin flow, rapid diffusion occurs, triggering self-assembly of nanoparticles (Figure 2-A). The flow rate ratio (FRR) between streams determines the extent of focusing, allowing precise control over particle characteristics. MHF also allows simultaneous encapsulation of hydrophilic drugs by adjusting the stream configurations.8,9
- Chaotic Advection Mixers:
These systems incorporate specially designed microchannels, such as herringbone patterns or serpentine paths, that disrupt laminar flow and induce rapid mixing through stretching and folding of fluid layers (Figure 2-B). This ensures that solvents and lipids mix quickly, initiating nanoparticle formation with improved uniformity. Chaotic advection combines the benefits of fast mixing with continuous-flow operation. 10–12. Explore the full potential of liposome production using this method for drug delivery.
- Vortex Focusing:
A hybrid of MHF and chaotic mixing, this approach uses a conical chamber where the lipid solution enters axially, and the buffer enters tangentially (Figure 2-C). The resulting spiral flow simultaneously focuses on the lipid stream and enhances mixing through rotational motion, enabling efficient nanoparticle formation in a single step.13,14
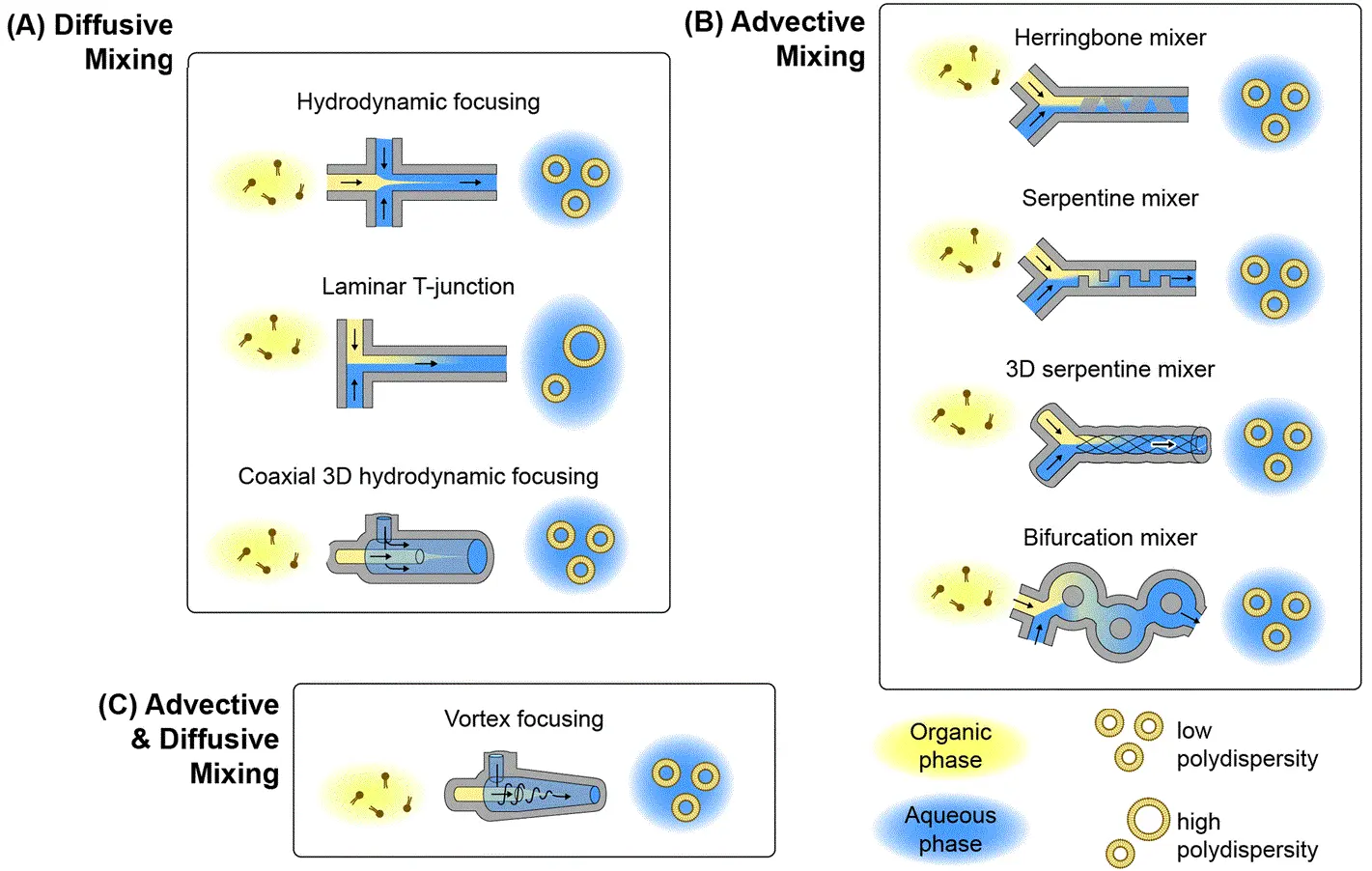
Figure 2: Microfluidic lipid nanoparticle production techniques
(From Mehraji, S. et al.; Lab Chip 2024, 24 (5), 1154–1174).
Explore more case studies of microfluidic techniques in LNP production:
2- Microfluidic Synthesis of Polymeric Nanoparticles:
Polymeric nanoparticles (PNPs) are increasingly used in drug delivery due to their versatile structure and the ability to encapsulate a wide range of therapeutic agents. These include hydrophilic and hydrophobic molecules, nucleic acids, and proteins. Unlike lipid-based nanoparticles, which are primarily lipid-based self-assemblies, PNPs are composed of polymeric materials such as poly(lactic-co-glycolic acid) (PLGA), polycaprolactone (PCL), and other biodegradable or biocompatible polymers. These polymers can form either matrix-type nanospheres or core-shell nanocapsules, providing flexibility in encapsulating different bioactive substances.9,15
The microfluidic synthesis of PNPs shares similarities with LNP production in the areas of precise control over particle size, size distribution, and encapsulation efficiency (Figure 3). Techniques for PNP synthesis include hydrodynamic focusing, nanoprecipitation, and coaxial flow systems, all of which are also employed in LNP synthesis. The primary difference lies in the material composition and the solvent systems used. 2,5,15,16
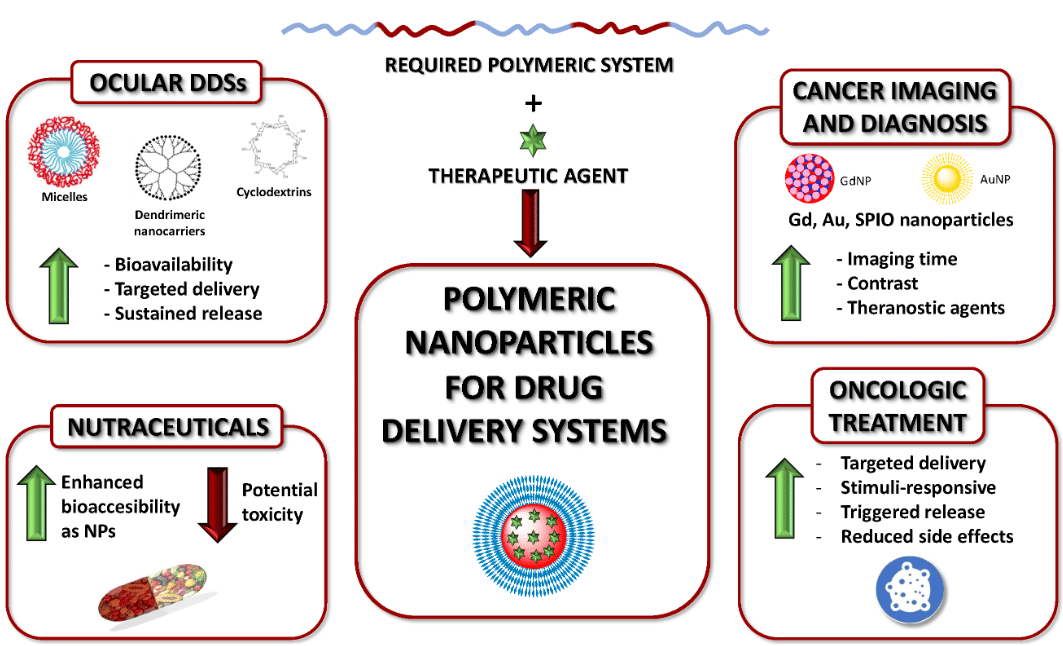
Figure 3: Polymeric nanoparticles for drug delivery
(From Begines, B. et al.; Nanomaterials 2020, 10 (7), 1403).
3- Microfluidic Production of Droplet-Based Microparticles
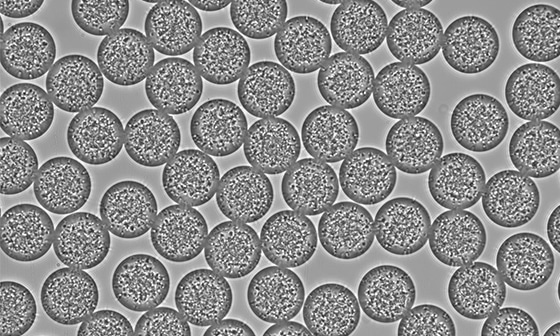
Microparticles made from biopolymers are emerging as important platforms for sustained drug delivery, cell therapy, and biomedical implants. Traditional batch production methods often suffer from poor size control and low reproducibility. Microfluidics enables the generation of uniform microparticles with precisely engineered properties, ensuring high-quality drug delivery systems.
This technique involves the generation of droplets within microchannels, where polymer solutions are encapsulated in immiscible carrier fluids. These droplets are then solidified through processes such as crosslinking, solvent evaporation, or polymerization. The geometry of the microchannels and the flow conditions can be finely tuned to control droplet size and formation frequency.5,9,17–19
These methods offer precision, reproducibility, and compatibility with various drug types and materials, making them an attractive choice for next-generation therapeutic systems.
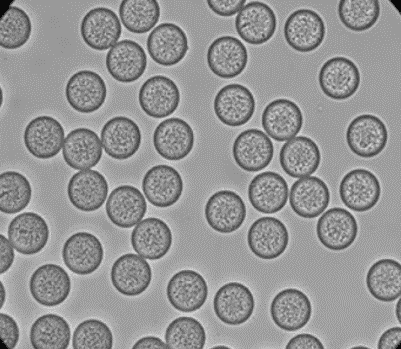
Figure 4: PLGA Microbeads encapsulating lysozyme (Produced by Secoya Technologies).
Table 1: Examples of drug encapsulation using microfluidics (Adapted from Parra Saldivar et al.; Front Biosci 2018, 10 (1), 74–91).
| Material Used | Geometry | Carrier Material | Drug | Application | Ref. |
|---|---|---|---|---|---|
| Glass | Co-flow | Human serum albumin, poly(lactic acid) | Doxorubicin | Hepatic cancer | 20 |
| PEEK and silica tube | T-junction | Poly(methyl acrylate), poly(acrylamide) | Ketoprofen, ranitidine | Suppression of gastric irritation effects | 21 |
| Silicon | Flow focusing | PLGA | Ciclosporin | Immunosuppressive therapy | 22 |
| PDMS | Co-flow with herringbone shape | Liposomes | Propofol | Anesthetic agent | 23 |
| Quartz Chip | Flow-focusing | Hyaluronic acid, ethylenediamine | Dexamethasone | Cell differentiation of mesenchymal stem cells | 24 |
| Glass | Co-flow and flow-focusing | Polycaprolactone, poly(vinyl alcohol), poly(ethylene glycol) | Bovine serum albumin | Protein therapy | 25 |
| PDMS | T-junction | Poly(ethylene glycol) diacrylate | 5-fluorouracil | Cancer therapy | 26 |
| PMMA | V-junction | Poly(methylsilsesquioxane) | Itraconazole | Antifungal drug for infections | 27 |
Explore more applications on droplet-based microfluidics:
- Microfluidic Application Notes
1-10 microns PLGA microsphere production using the RayDrop
Read more - Microfluidic Application Notes
Alginate Microcapsule Synthesis
Read more
Free content for better understanding on droplet production methods
Based Microneedle Systems for Carrier-Free Drug Delivery
Microfluidic technology enables the advancement of carrier-free drug delivery systems. By leveraging the precision and control provided by microfluidic platforms, drugs can be delivered directly to target sites, minimizing the need for conventional drug carriers. This results in a more efficient delivery system with better bioavailability and targeted release. Among the most promising components in this approach are micro-needles, which are highly integrated into microfluidic devices to optimize the delivery of therapeutics.2,28,29
Micro-Needles (MNs): These are devices that utilize arrays of microsized-needles, small, precise structures designed to penetrate the skin or other tissues for targeted drug delivery. These microneedles enable painless, minimally invasive drug administration, bypassing the need for traditional injections and oral delivery systems. The control offered by microfluidics ensures localized and controlled release of the drug, which can enhance the bioavailability and targeted action of therapeutics, particularly those that may degrade in the digestive system.
Types of Micro-Needles:
- Solid Micro-Needles: These are designed to create micro-channels in the skin, after which drugs can be applied topically, allowing them to diffuse passively through the skin. Solid MNs are typically made from materials such as (Silicon, Metals, Polymers).30
- Dissolving Micro-Needles: Composed of biodegradable materials, these MNs dissolve upon insertion into the skin, releasing the drug directly into the targeted tissue. Materials used for dissolving MNs include (Polyvinyl Alcohol (PVA), Polyvinylpyrrolidone (PVP), Polylactic Acid (PLA)).31
- Hydrogel Micro-Needles: These are made from swelling polymer materials and release drugs via the expansion of the hydrogel upon insertion into the skin. Materials for hydrogel MNs include (Polyethylene Glycol (PEG), Polyacrylamide (PAAm), Chitosan).32
- Hollow Micro-Needles: These are designed with a hollow core that allows for the direct infusion of drugs into the body through the needle. Hollow MNs are typically made from materials such as (Glass, Silicon, Metals, Polymers).33,34
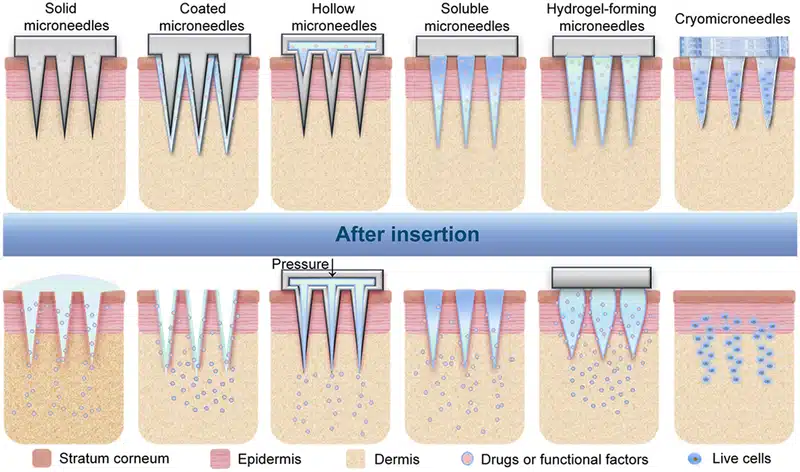
Figure 5: Types of microneedles and their corresponding drug delivery mechanisms
(From Zhang, Y. er al.; Exploration 2023, 3 (1), 20210170).
In Vitro Models for the Evaluation of Drug Delivery
To bridge the gap between conventional in vitro models and the complexity of human physiology, organ-on-a-chip (OOC) platforms have emerged as powerful tools in the evaluation of drug delivery systems. These models closely replicate the dynamic architecture and function of human tissues, offering insights into drug transport, absorption, and therapeutic response. As such, microfluidics in drug delivery has become an emerging technology for studying how nanocarriers interact with traverse biological barriers.35,36
Once a drug carrier enters the body, its journey to the target site is impeded by several key physiological barriers. Accurately modeling and understanding these obstacles is critical to the development of effective and safe drug delivery systems. Microfluidic OOC platforms offer a highly controlled environment where researchers can simulate and analyze these barriers in real time.
Key biological barriers include:
- Blood–Brain Barrier (BBB): This tightly regulated interface restricts the entry of most therapeutic agents into the brain. Microfluidic BBB-on-a-chip models co-culture endothelial cells, astrocytes, and pericytes under controlled shear flow to mimic the selective permeability and tight junctions characteristic of the human BBB. These systems are vital for evaluating brain-targeted drug delivery systems. 37
- Mucosal Diffusion Barrier: Present in areas such as the gastrointestinal, respiratory, and reproductive tracts, this consists of a dense mucus layer that traps and excludes foreign particles. Microfluidic models replicate mucus viscosity and secretion dynamics, allowing real-time observation of nanoparticle diffusion, penetration, and retention for oral and pulmonary drug delivery.38
- Cellular Permeability Barrier: Composed of epithelial or endothelial cell monolayers with tight junctions, this barrier controls both transcellular and paracellular transport. On-chip systems can simulate cellular architecture and mechanical stimuli, enabling studies of nanoparticle uptake, receptor-mediated transport, and barrier integrity modulation.39
- Biochemical Barrier: Enzymes and pH variations, particularly in the gut and lysosomal environments, can degrade or inactivate drugs before they reach their target. Microfluidic biochemical models simulate these conditions to test the stability and protective efficacy of nanocarriers under physiologically relevant stresses.2
The integration of these barriers into microfluidic organ-on-a-chip platforms enables more predictive and human-relevant preclinical evaluation.
Devices such as, (gut-on-a-chip, vessel-on-a-chip, blood–brain barrier-on-a-chip have demonstrated the potential of precision drug delivery using microfluidics.
As microfluidics in drug delivery continues to evolve, these platforms can be used in nanomedicine for evaluating pharmacokinetics, pharmacodynamics, and therapeutic index in a more human-relevant manner with reduced dependence on animal models.
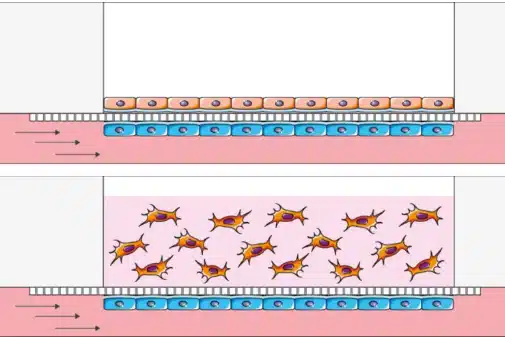
Figure 6: Creation of an endothelium-epithelium barrier for air liquid interface cell culture in 2D or 3D
(From BeOnChip).
An example in ocular drug delivery involves the application of microfluidic systems to replicate dynamic intraocular environments. By integrating flow control and temperature regulation, these platforms allow for the simulation of physiological conditions, such as intraocular pressure fluctuations and eye motion. This enables precise assessment of how these variables affect drug clearance and retention. The combination of automated sensing and synchronized flow across multiple in vitro models improves reproducibility and throughput in formulation screening, supporting more predictive and scalable testing for intraocular drug delivery.40
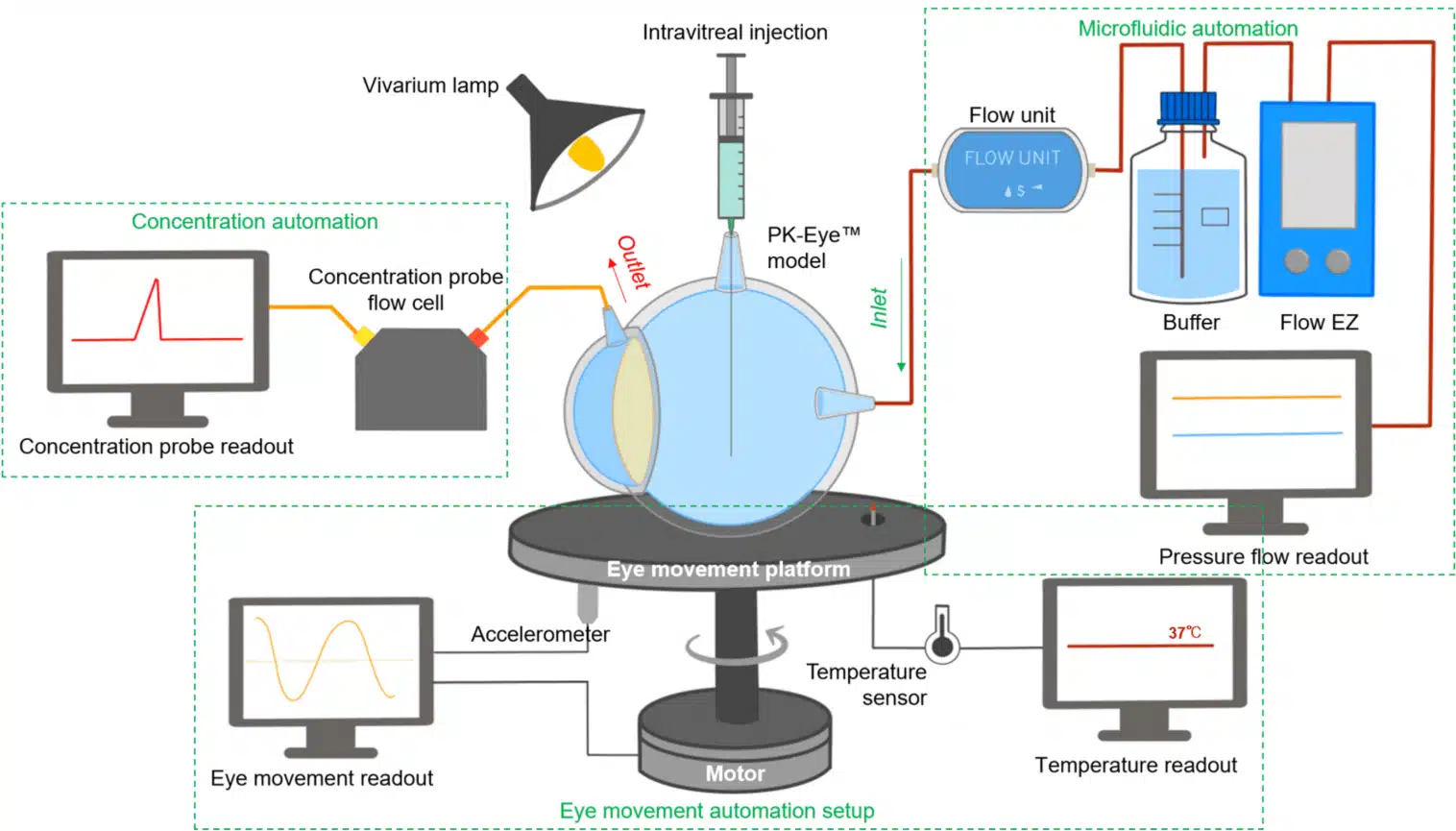
Figure 7: In vitro microfluidic monitoring platform for ocular drug delivery
(Awwad, S. el al.; Pharmaceutics 2023, 15 (5), 1444).
Microfluidics for Drug and Protein Crystallization in Pharmaceutical Research
Microfluidic systems are also transforming protein crystallization, a key process in drug development. Traditionally requiring large sample volumes, microfluidic platforms allow for the screening of crystallization conditions with minimal amounts of protein and reagents. By simulating various conditions like pH, temperature, and salt concentrations, these devices enable faster, more efficient crystallization.
Recent innovations, such as centrifuge-based microfluidic devices and semi-contact dispensing methods, have enhanced high-throughput screening and precision, reducing costs and accelerating drug development. This advancement plays a crucial role in creating high-quality protein crystals for structural analysis, aiding in more targeted drug delivery.41–43

An all-in-one platform for continuous generation of UV-cured core-shell microcapsules
Read More
Webinar: Controlled UV-crosslinked microcapsule production using microfluidic technology
Read More
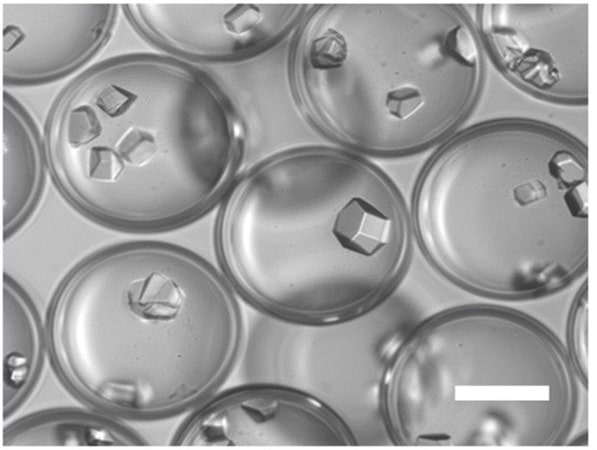
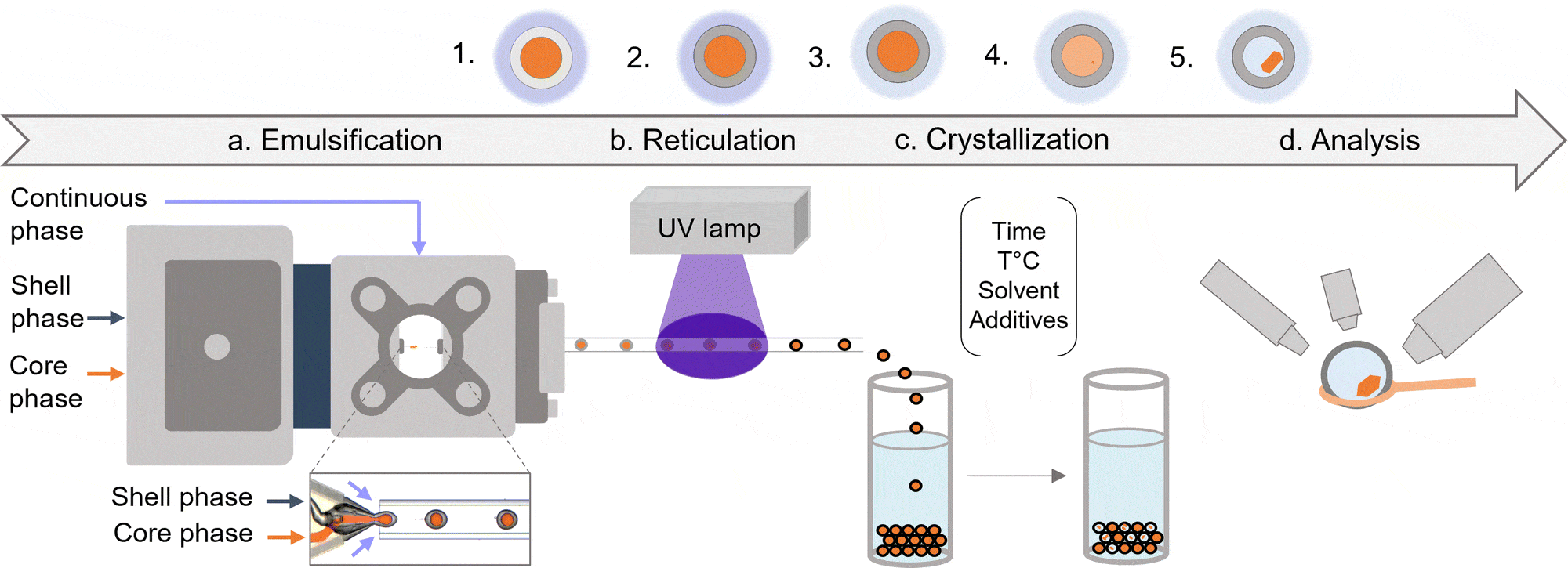

Figure 8: (Left)Schematic representation of an example of crystallization process from initial droplets. (Right) Lysozymes crystals inside microcapsules
(From Mettler, M. et al.; Chem. Commun. 59, 12739–12742 (2023)).
Conclusion
In this review, we highlighted the advancements in drug delivery systems enabled by microfluidics, which enhance bioavailability, drug efficiency, and nanoparticle performance. Despite progress, challenges remain in scaling these systems for clinical use, with the need for improved parallelization and simplified fabrication. Integrating microfluidics with organ-on-a-chip technology offers promising solutions for more accurate preclinical testing and personalized medicine. With continued interdisciplinary collaboration, microfluidics has the potential to transform further drug delivery and therapeutic applications.
👉 Ready to improve your drug delivery workflows? Contact our experts or explore our microfluidic pressure control systems.
Related Products
Expertises and resources
-
Expert Reviews: Basics of Microfluidics 10 Tips for Reliable Droplet Generation Read more
-
Microfluidics Case Studies Drug-Loaded Liposome Preparation Using Microfluidics Read more
-
Microfluidics Case Studies Real-Time Monitoring Platform for Ocular Drug Delivery, Integrating Fluigent’s Flow EZ Read more
-
Microfluidic Application Notes 1-10 microns PLGA microsphere production using the RayDrop Read more
-
Microfluidics Article Reviews Solid lipid nanoparticles for biologics and drug encapsulation Read more
-
Microfluidic Application Notes Alginate Microcapsule Synthesis Read more
-
Microfluidic Application Notes Encapsulation of multiple emulsions in a single droplet Read more
-
Microfluidics Article Reviews Human Blood Brain Barrier (BBB) permeability -on-chip assessment Read more
-
Microfluidics White Papers A review of Organ on Chip Technology – A White Paper Read more
-
Microfluidics Article Reviews Microfluidic technology for engineered nanoparticles in nanomedicine Read more
-
Microfluidic Application Notes Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions Read more
-
Microfluidic Application Notes PLGA microcapsules synthesis Read more
-
Microfluidics White Papers Droplet-based Microfluidics – A Complete Guide Read more
-
Microfluidic Application Notes Liposome Nanoparticle Synthesis Read more
-
Expert Reviews: Basics of Microfluidics Flow Control Technologies: Comparison between peristaltic, syringe and pressure pumps for microfluidic applications Read more
References
1. Park, K. Controlled drug delivery systems: Past forward and future back. J. Controlled Release 190, 3–8 (2014).
2. Ma, Z., Li, B., Peng, J. & Gao, D. Recent Development of Drug Delivery Systems through Microfluidics: From Synthesis to Evaluation. Pharmaceutics 14, 434 (2022).
3. Li, C. et al. Recent progress in drug delivery. Acta Pharm. Sin. B 9, 1145–1162 (2019).
4. Jaradat, E., Weaver, E., Meziane, A. & Lamprou, D. A. Microfluidics Technology for the Design and Formulation of Nanomedicines. Nanomaterials 11, 3440 (2021).
5. Parra Saldivar, R. Microfluidics technology for drug delivery A review. Front. Biosci. 10, 74–91 (2018).
6. Zhang, H. et al. Microfluidics for nano-drug delivery systems: From fundamentals to industrialization. Acta Pharm. Sin. B 13, 3277–3299 (2023).
7. Mehraji, S. & DeVoe, D. L. Microfluidic synthesis of lipid-based nanoparticles for drug delivery: recent advances and opportunities. Lab. Chip 24, 1154–1174 (2024).
8. Jahn, A., Vreeland, W. N., Gaitan, M. & Locascio, L. E. Controlled Vesicle Self-Assembly in Microfluidic Channels with Hydrodynamic Focusing. J. Am. Chem. Soc. 126, 2674–2675 (2004).
9. Jahn, A. et al. Preparation of nanoparticles by continuous-flow microfluidics. J. Nanoparticle Res. 10, 925–934 (2008).
10. Suh, Y. K. & Kang, S. A Review on Mixing in Microfluidics. Micromachines 1, 82–111 (2010).
11. Nguyen, N.-T. & Wu, Z. Micromixers—a review. J. Micromechanics Microengineering 15, R1–R16 (2005).
12. Saorin, A. et al. Microfluidic production of amiodarone loaded nanoparticles and application in drug repositioning in ovarian cancer. Sci. Rep. 14, 6280 (2024).
13. Markwalter, C. E. & Prud’homme, R. K. Design of a Small-Scale Multi-Inlet Vortex Mixer for Scalable Nanoparticle Production and Application to the Encapsulation of Biologics by Inverse Flash NanoPrecipitation. J. Pharm. Sci. 107, 2465–2471 (2018).
14. Liu, Y., Cheng, C., Liu, Y., Prud’homme, R. K. & Fox, R. O. Mixing in a multi-inlet vortex mixer (MIVM) for flash nano-precipitation. Chem. Eng. Sci. 63, 2829–2842 (2008).
15. Leung, M. H. M. & Shen, A. Q. Microfluidic Assisted Nanoprecipitation of PLGA Nanoparticles for Curcumin Delivery to Leukemia Jurkat Cells. Langmuir 34, 3961–3970 (2018).
16. Begines, B. et al. Polymeric Nanoparticles for Drug Delivery: Recent Developments and Future Prospects. Nanomaterials 10, 1403 (2020).
17. Onan, D. et al. Microfluidics Based Particle and Droplet Generation for Gene and Drug Delivery Approaches. J. Biomed. Mater. Res. B Appl. Biomater. 113, e35530 (2025).
18. Hallan, S. S., Kaur, P., Kaur, V., Mishra, N. & Vaidya, B. Lipid polymer hybrid as emerging tool in nanocarriers for oral drug delivery. Artif. Cells Nanomedicine Biotechnol. 44, 334–349 (2016).
19. Mukherjee, A. et al. Lipid–polymer hybrid nanoparticles as a next-generation drug delivery platform: state of the art, emerging technologies, and perspectives. Int. J. Nanomedicine Volume 14, 1937–1952 (2019).
20. Kim, M.-K., Kim, M. A., Jenjob, R., Lee, D.-H. & Yang, S.-G. Capillary microfluidics-derived doxorubicin-containing human serum albumin microbeads for transarterial chemoembolization of hepatic cancer. Mater. Sci. Eng. C 62, 391–397 (2016).
21. Khan, I. U. et al. Microfluidic conceived pH sensitive core–shell particles for dual drug delivery. Int. J. Pharm. 478, 78–87 (2015).
22. Keohane, K., Brennan, D., Galvin, P. & Griffin, B. T. Silicon microfluidic flow focusing devices for the production of size-controlled PLGA based drug loaded microparticles. Int. J. Pharm. 467, 60–69 (2014).
23. Kastner, E., Verma, V., Lowry, D. & Perrie, Y. Microfluidic-controlled manufacture of liposomes for the solubilisation of a poorly water soluble drug. Int. J. Pharm. 485, 122–130 (2015).
24. Agnello, S. et al. Microfluidic production of hyaluronic acid derivative microfibers to control drug release. Mater. Lett. 182, 309–313 (2016).
25. Pessi, J. et al. Microfluidics-assisted engineering of polymeric microcapsules with high encapsulation efficiency for protein drug delivery. Int. J. Pharm. 472, 82–87 (2014).
26. Xue, P., Wu, Y., Menon, N. V. & Kang, Y. Microfluidic synthesis of monodisperse PEGDA microbeads for sustained release of 5-fluorouracil. Microfluid. Nanofluidics 18, 333–342 (2015).
27. Kucuk, I., Ahmad, Z., Edirisinghe, M. & Orlu-Gul, M. Utilization of microfluidic V-junction device to prepare surface itraconazole adsorbed nanospheres. Int. J. Pharm. 472, 339–346 (2014).
28. Riahi, R. et al. Microfluidics for advanced drug delivery systems. Curr. Opin. Chem. Eng. 7, 101–112 (2015).
29. Bilal, M. et al. Microneedles in Smart Drug Delivery. Adv. Wound Care 10, 204–219 (2021).
30. Pradeep Narayanan, S. & Raghavan, S. Solid silicon microneedles for drug delivery applications. Int. J. Adv. Manuf. Technol. 93, 407–422 (2017).
31. Ahmad, N. N., Ghazali, N. N. N. & Wong, Y. H. Concept Design of Transdermal Microneedles for Diagnosis and Drug Delivery: A Review. Adv. Eng. Mater. 23, 2100503 (2021).
32. Chen, S. et al. Microneedle‐Array Patch Fabricated with Enzyme‐Free Polymeric Components Capable of On‐Demand Insulin Delivery. Adv. Funct. Mater. 29, 1807369 (2019).
33. Trautmann, A., Roth, G.-L., Nujiqi, B., Walther, T. & Hellmann, R. Towards a versatile point-of-care system combining femtosecond laser generated microfluidic channels and direct laser written microneedle arrays. Microsyst. Nanoeng. 5, 6 (2019).
34. Zhang, Y. et al. Microneedle system for tissue engineering and regenerative medicine. Exploration 3, 20210170 (2023).
35. Bhatia, S. N. & Ingber, D. E. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
36. Van Der Helm, M. W. et al. Direct quantification of transendothelial electrical resistance in organs-on-chips. Biosens. Bioelectron. 85, 924–929 (2016).
37. Begley, D. J. Delivery of therapeutic agents to the central nervous system: the problems and the possibilities. Pharmacol. Ther. 104, 29–45 (2004).
38. Jia, Z., Guo, Z., Yang, C.-T., Prestidge, C. & Thierry, B. “Mucus-on-Chip”: A new tool to study the dynamic penetration of nanoparticulate drug carriers into mucus. Int. J. Pharm. 598, 120391 (2021).
39. Waheed, S. et al. Engineering nano-drug biointerface to overcome biological barriers toward precision drug delivery. J. Nanobiotechnology 20, 395 (2022).
40. Awwad, S. et al. Real-Time Monitoring Platform for Ocular Drug Delivery. Pharmaceutics 15, 1444 (2023).
41. Mettler, M., Dewandre, A., Tumanov, N., Wouters, J. & Septavaux, J. Single crystal formation in core–shell capsules. Chem. Commun. 59, 12739–12742 (2023).
42. Wang, L. et al. A centrifugal microfluidic device for screening protein crystallization conditions by vapor diffusion. Sens. Actuators B Chem. 219, 105–111 (2015).
43. Zhu, Y. et al. Nanoliter-Scale Protein Crystallization and Screening with a Microfluidic Droplet Robot. Sci. Rep. 4, 5046 (2014).
A University College London (UCL) Paper
Paper: Awwad, S.; Ibeanu, N.; Liu, T.; Velentza-Almpani, A.; Chouhan, N.; Vlatakis, S.; Khaw, P. T.; Brocchini, S.; Bouremel, Y. Real-Time Monitoring Platform for Ocular Drug Delivery. Pharmaceutics 2023, 15 (5), 1444. https://doi.org/10.3390/pharmaceutics15051444
Part of the Faculty of Brain Sciences, the UCL Institute of Ophthalmology holds the top global ranking for ophthalmology studies (per CWUR Rankings by Subject 2017). Collaborating with Moorfields Eye Hospital, they form the largest co-located hub for eye research, education, and care worldwide. Through groundbreaking research in ophthalmology and eye health, their interdisciplinary approach unites scientists, clinicians, and patients, leading to tangible improvements in people’s lives.
In collaboration with UCL, Optceutics Ltd. manufactures advanced in vitro models of the human eye (PK-Eye™) that accelerate the development of longer-acting intraocular medicines. Optceutics Ltd. offers fee-for-service research and strategic collaborations, specializing in preclinical ophthalmic formulation optimization.



Challenges in Treating Posterior Eye Diseases
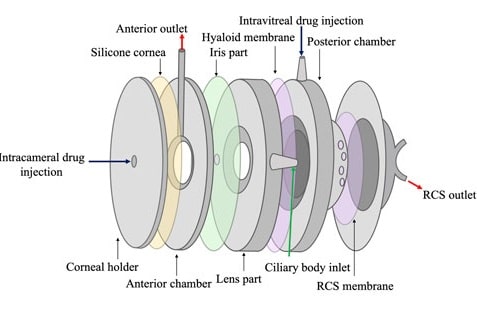
The posterior segment of the eye, where diseases like age-related macular degeneration, glaucoma, and diabetic retinopathy occur, poses significant challenges for treatment due to its complexity. These ocular diseases, which can lead to blindness, affect millions worldwide and are becoming more prevailing with the increase in the aging population.[1] To treat them, therapeutic antibodies and proteins have transformed the management of such conditions, often requiring direct injections into the eye for effective ocular drug delivery (figure 1). However, frequent intravitreal injections are burdensome for patients and healthcare systems, prompting the need for improved drug formulations to prolong efficacy and reduce risks. [2,3] Therefore, formulation strategies are essential to address these challenges, necessitating either progress preclinical evaluation in animal models or in vitro models.
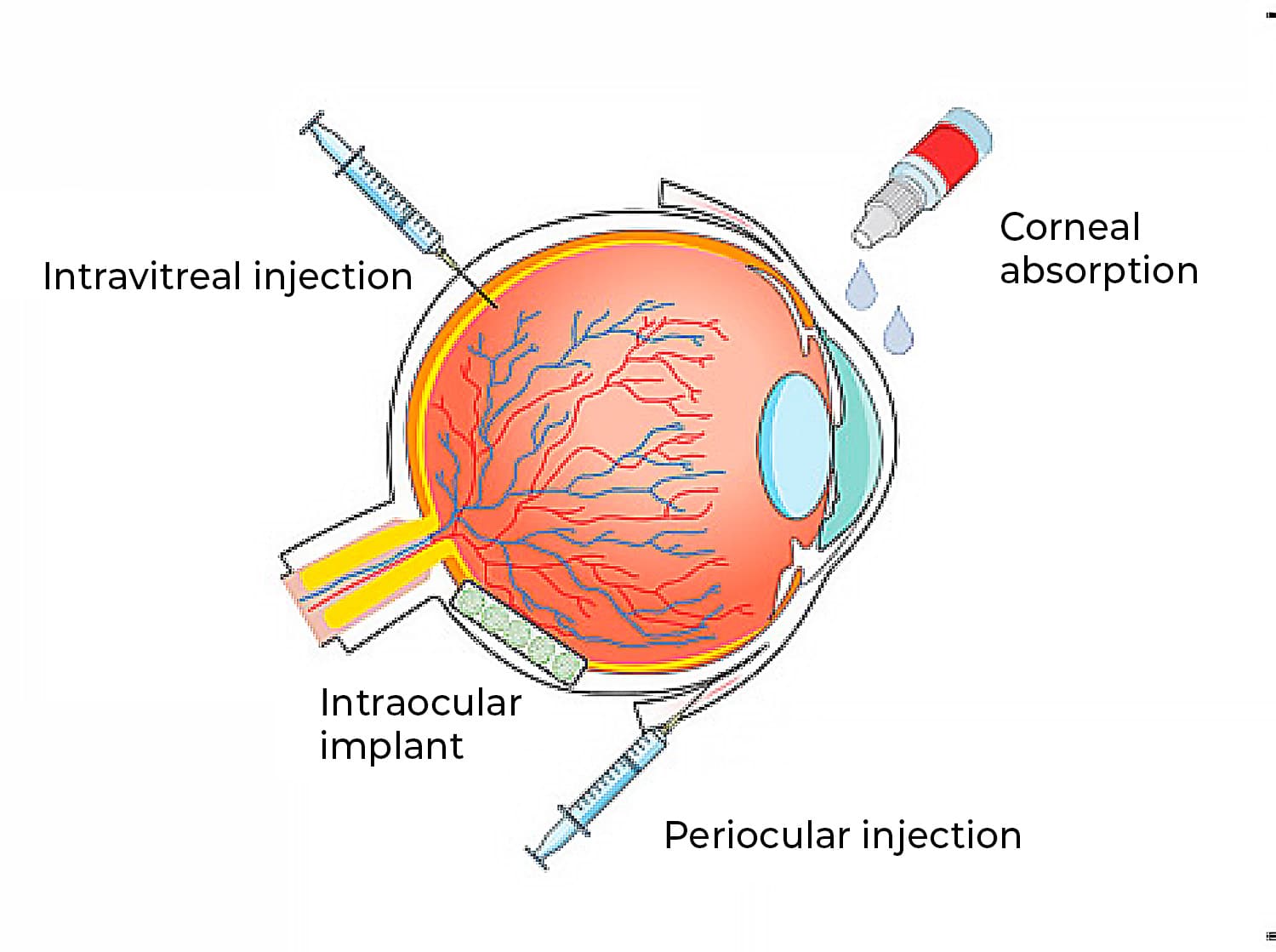
Figure 1: Ocular routes for drug delivery.[4]
How Can in Vitro Models Overcome Limitations in Ocular Drug Delivery Compared to Animal Models?
Classical in-animal models present several limitations in research and drug discovery, including anatomical differences between animal and human eyes. In addition, the formation of anti-drug antibodies complicates the testing of biologics. Ocular tolerability and high costs associated with animal models further highlight the need for alternative testing techniques.[5]
In vitro models offer a promising solution, serving as alternatives or supplements to animal testing during preclinical development. These models not only reduce reliance on animal studies but also help validate dissolution specifications and demonstrate the bioequivalence of active pharmaceutical ingredients. They have been designed for various aims such as assessing protein stability in simulated vitreous fluids, understanding the impact of eye movements, and evaluating drug release/clearance times (figure 2). [6]
Despite significant progress in in vitro ocular formulation testing for posterior eye diseases, there is currently no approved model specifically designed to assess intraocular pharmacokinetics and still lack adequate representation of eye dynamics, such as flow rate and compartmentalization, crucial for understanding drug kinetics and stability within the eye.
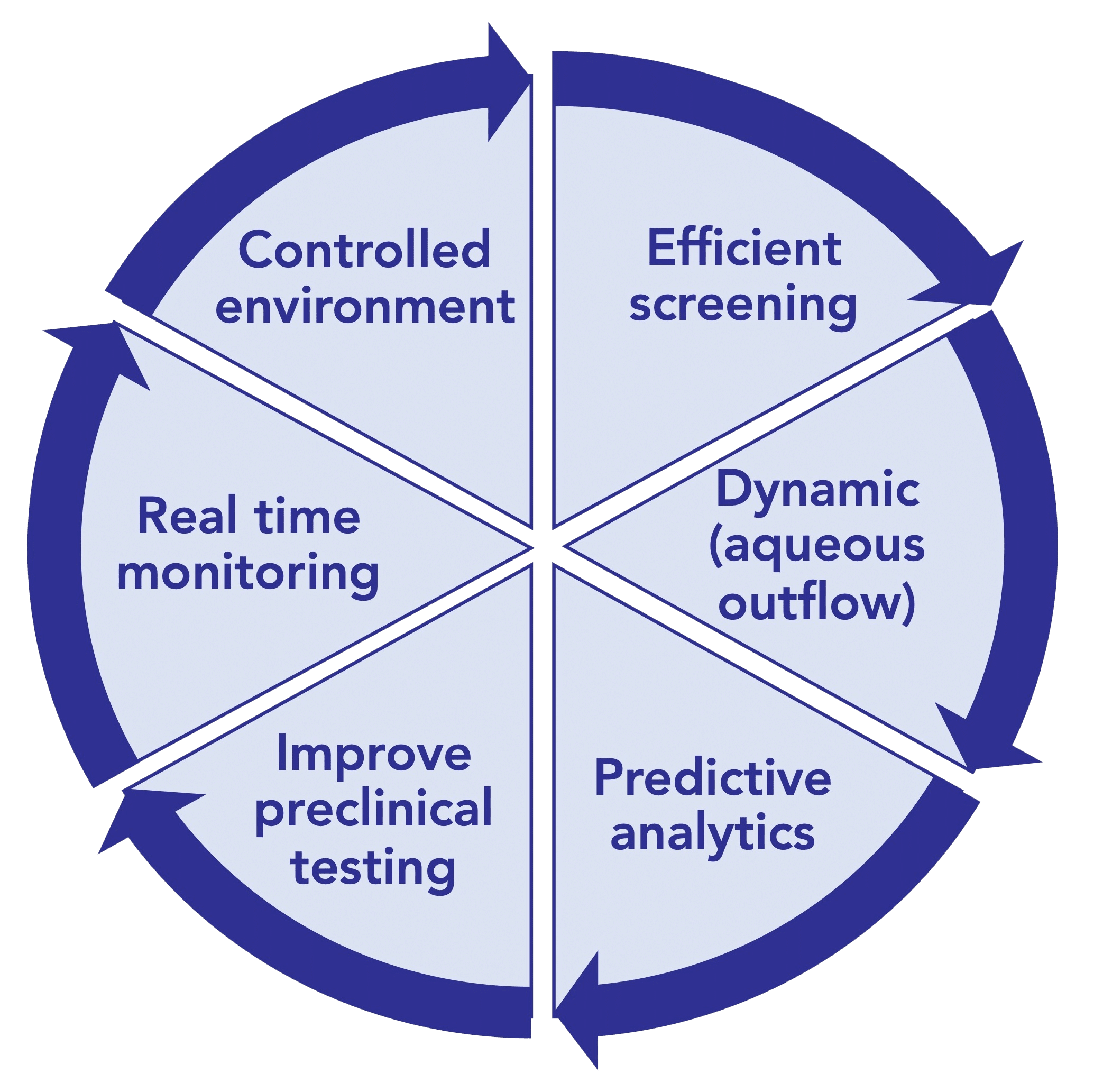
Figure 2: Ultimate goals for In-vitro eye models to evaluate ocular drug delivery.[7]
PK-Eye™ Model: Example of Real-Time Monitoring Platform for Ocular Drug Development.
The PK-Eye™ model, designed and manufactured by Optceutics Ltd., emerged as a robust and user-friendly tool, addressing the limitations of in vivo models for developing long-acting intraocular medicines. Combining this in vitro model with real-time monitoring systems will facilitate more efficient preclinical testing by increasing data collection while minimizing manual intervention.[8]
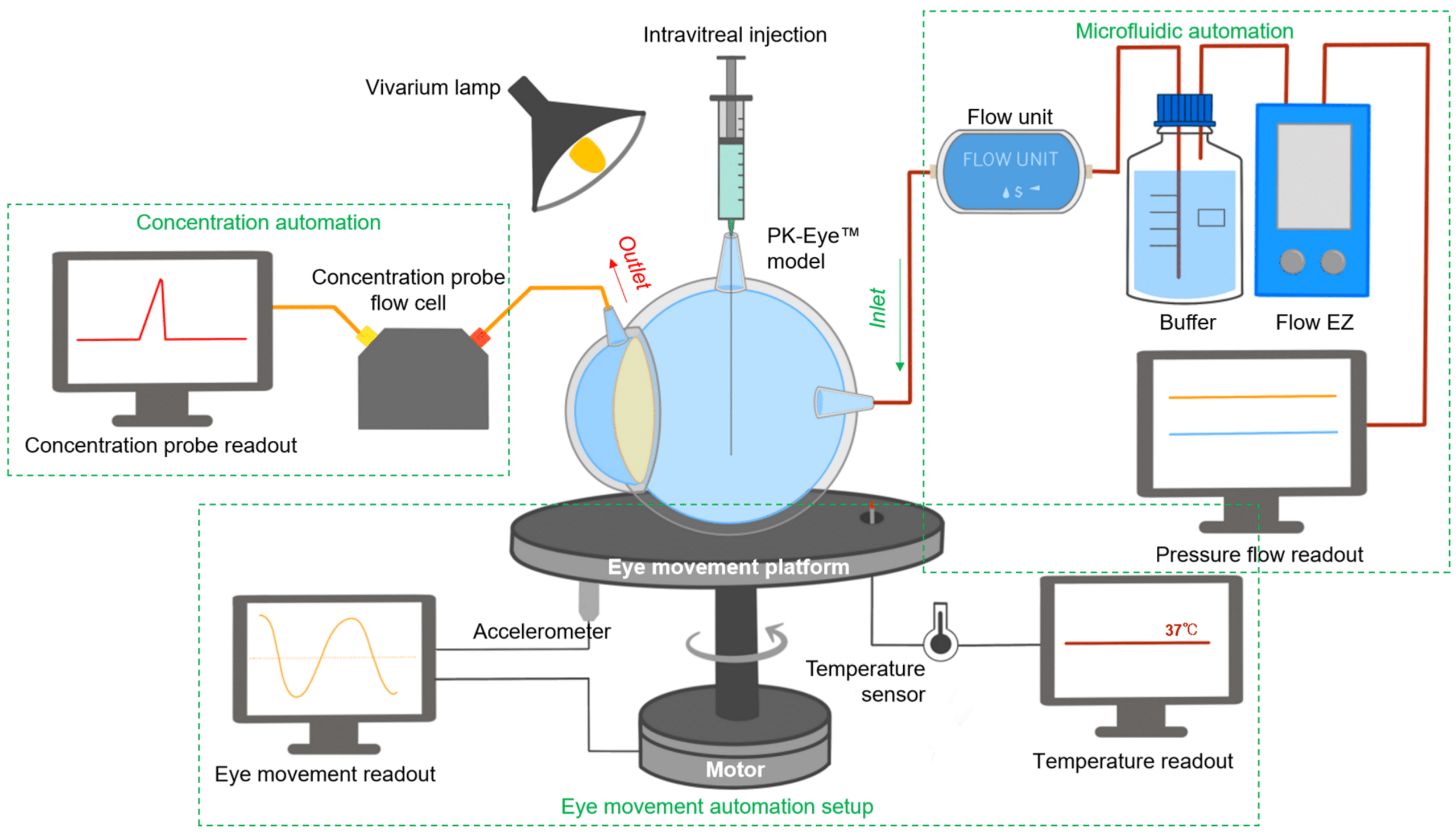
In a paper published in Pharmaceutics (2023), a newly designed real-time monitoring platform was scaled and automated for ocular drug delivery. This platform aims to provide proof-of-concept for a fully automated and optimized PK-Eye™ model testing setup, to accelerate intraocular drug development by streamlining sample analysis and improving accuracy (figure 3).
Key features of the platform included monitoring flow, temperature, eye movements, and concentration evaluation of labeled protein molecules (figure 4).
How to Scale up Real-Time Monitoring
Each PK-Eye™ model was connected to a FlowEZ microfluidic controller, with buffer (PBS, pH 7.4). These controllers were linked to an FLPG plus 2-bar pressure source. The pressure and flow were managed via the Fluigent A-i-O program (previous version of Oxygen). Scalability was tested with configurations running 1× LineUp Flow EZ controlling 1–6 models (figure 5). Experiments ran at a fixed flow rate of 2 μL/min at 37°C, with a 48-hour equilibration period before data recording. The flow rates were recorded with flow units S.
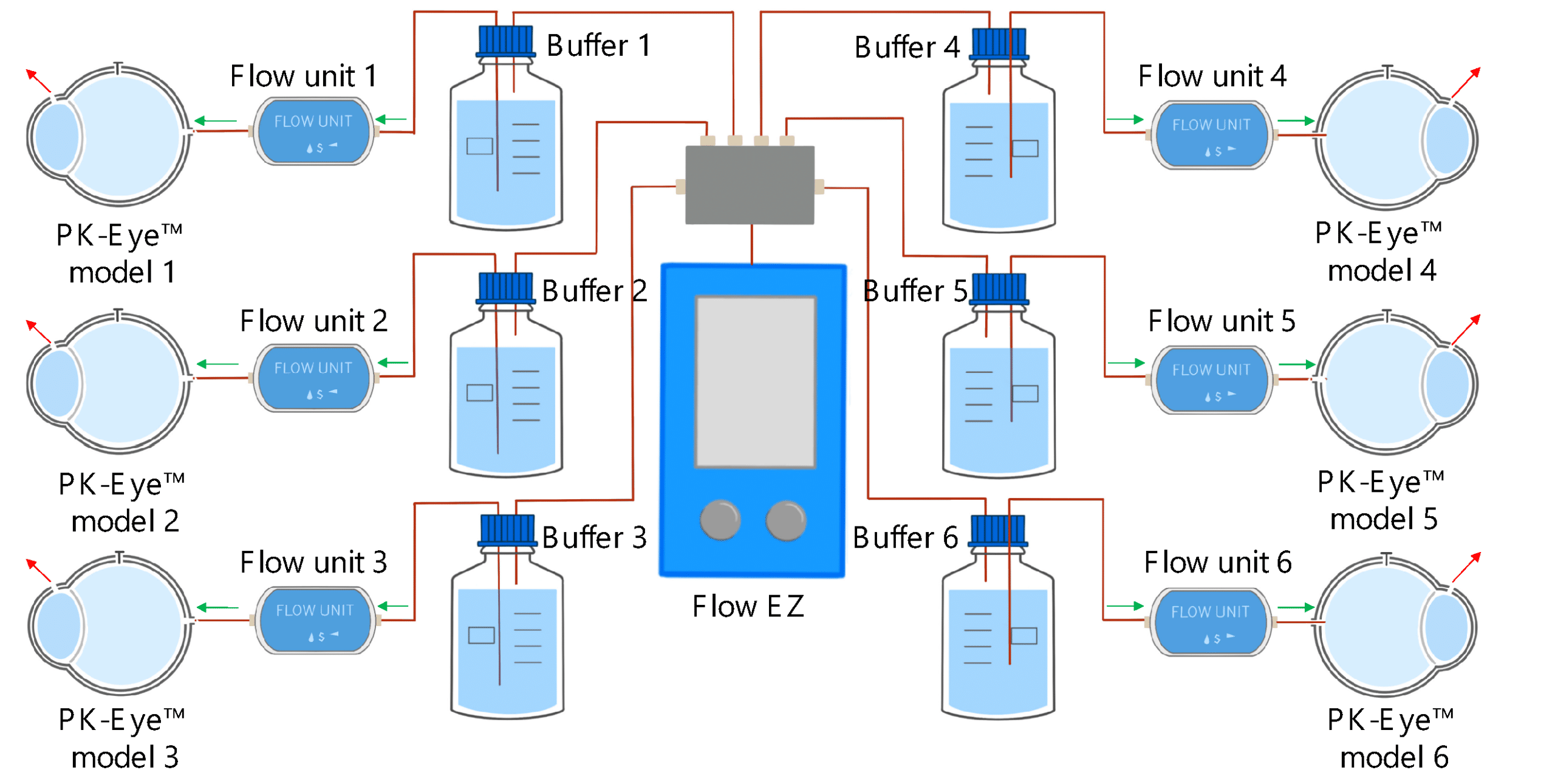
Figure 5: Experimental setup of flow rate control with 1:6 ratios using PK Eye models in PBS at RT with a pressure control of for 24h. [7]
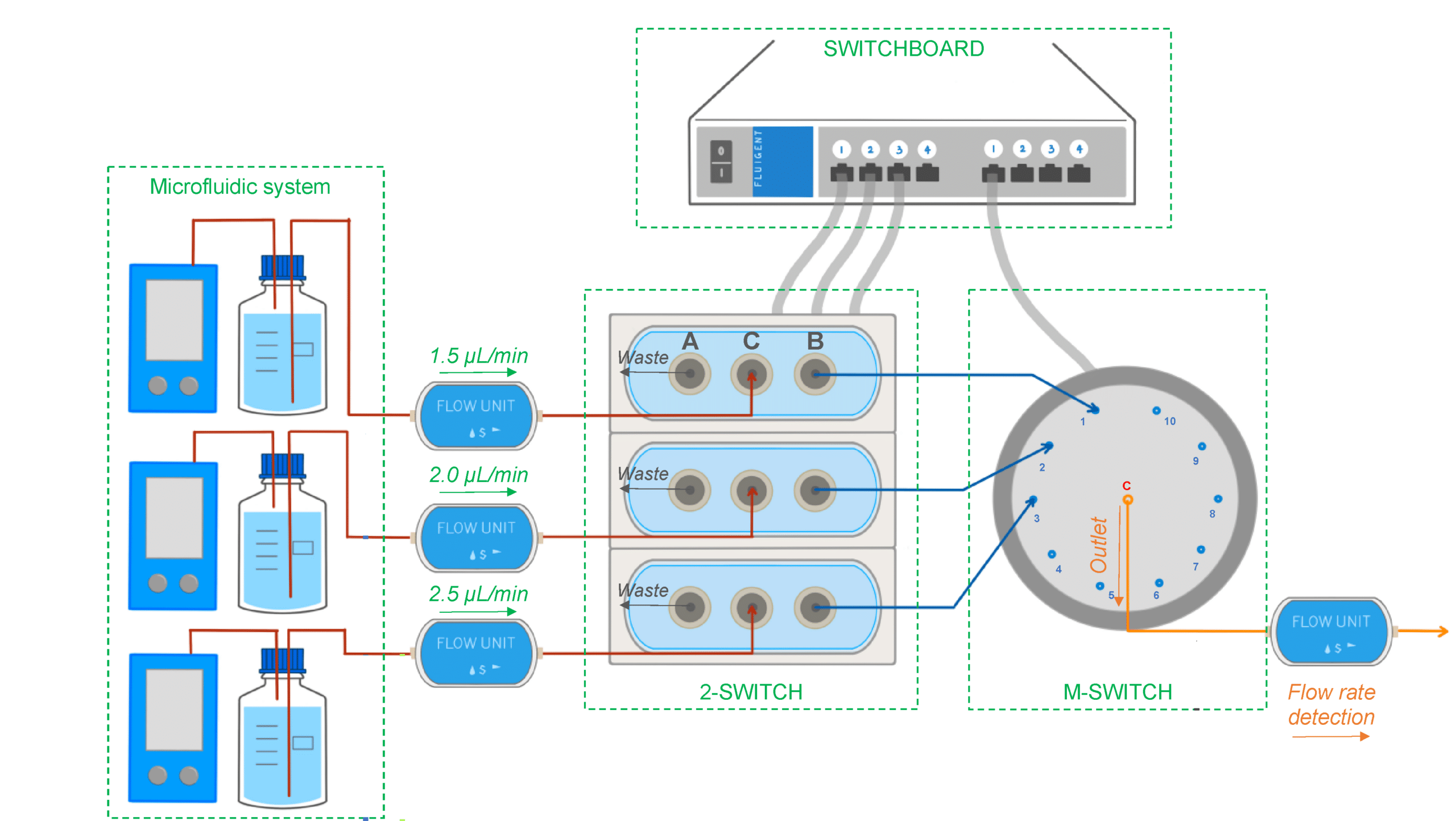
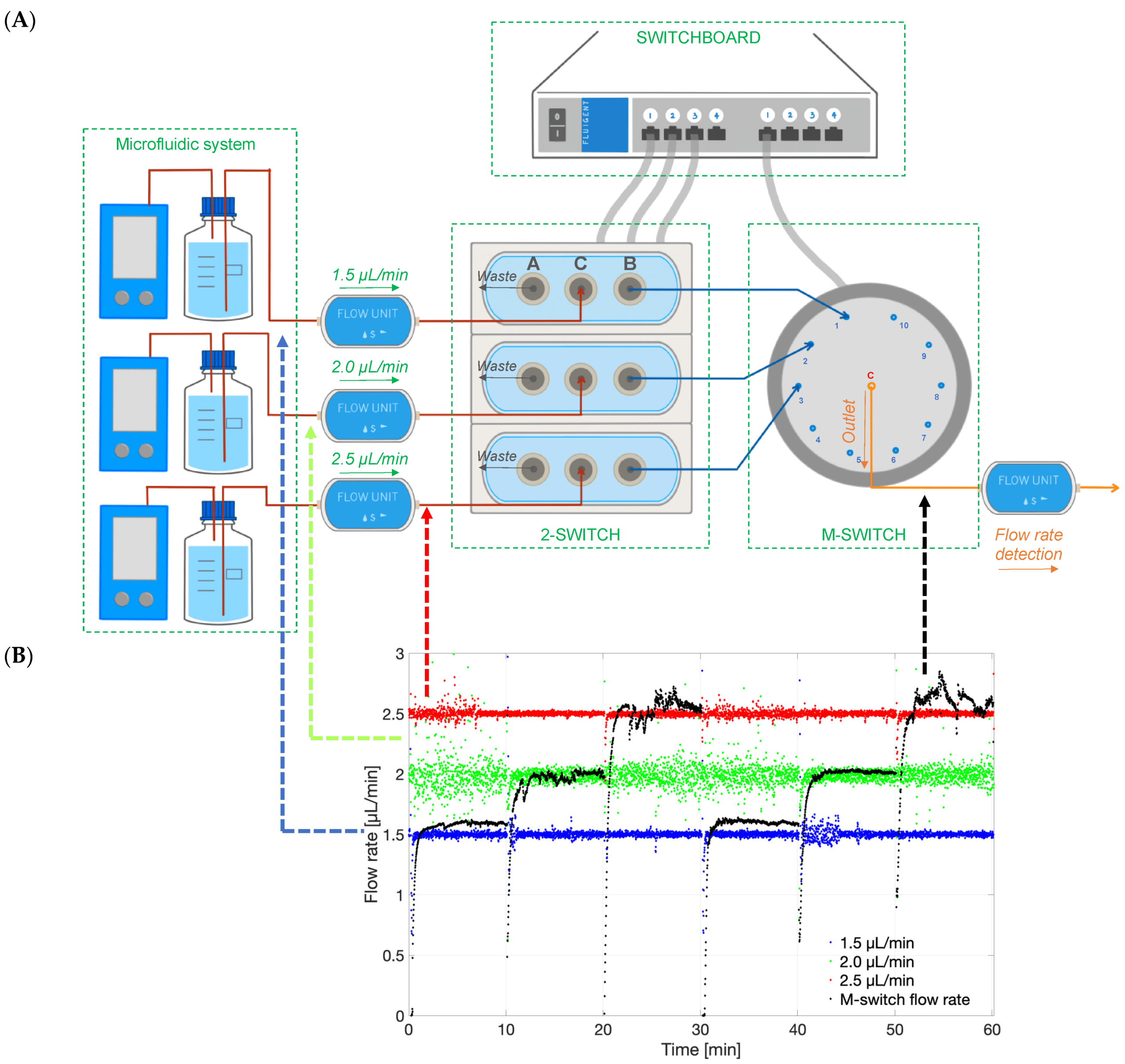
Figure 6: Experimental setup used to select different flow lines to be analyzed by the concentration detection unit to monitor multiple PK-Eye™ models.[7]
To enable real-time monitoring of drug release profiles across multiple models or fluid inlets, microfluidic valves from Fluigent were integrated into the system. It included three 3-port/2-way microfluidic valves (2-SWITCH), an 11-port/10-position rotary valve (M-SWITCH), and a SWITCHBOARD (Previous version of SWITCH EZ) (figure 6). This platform facilitated the selective collection of outflows from different models or inlets into the concentration detector, allowing for varied release profile readouts.
Next, the integration of this platform with a concentration probe was demonstrated. A PK-Eye™ model with a membrane was connected to the 2-SWITCH, allowing switching between fluid outlets. PBS buffer was pumped through the model while the outflow was alternated between the model and a PBS reservoir. Alexa albumin was injected into the model, and the system toggled between the model and reservoir every 2 hours for real-time detection by the concentration probe.
Proof-of-Concept: Unlock Stability and Scalability Using Flow Controllers vs Peristaltic Pumps
Traditional syringe or peristaltic pumps are commonly used in pharmaceutical labs to impose a constant flow rate, but they often result in cyclical flow rate variations. Fluigent’s pressure-driven controller (FlowEZ) was introduced in this work to allow large-scale experiments, offering smooth flow rates with minimal variation and rapid response to pressure or flow rate changes. Troubleshooting was also easier with this system compared to peristaltic pumps (table 1).
Table 1: Comparison between the PK-Eye platform based on a peristaltic pump system or a flow controller system.
| Model-Platform | Flow based on peristaltic pump (1st Generation PK-Eye™) | Flow based on pressure-driven controllers (Latest PK-Eye™ model) |
|---|---|---|
| Pressure-flow Monitoring | No | Yes (with graph readout) |
| Simultaneous models testing (n) | 8 | 48 |
| Circadian rhythm | No | Yes (with graph readout) |
| Simulated vitreous fluids leakage monitoring | No | Yes (with graph readout) |
| Eye movement monitoring | No | Yes, eye movements shown (with graph readout) |
| Temperature monitoring | Thermometer | Temperature sensors |
| Concentration readout | Manual sampling and HPLC analysis | Manual sampling and HPLC analysis, and use of concentration probe setup |

“A pressure-controlled flow system, as opposed to syringe pumps or peristaltic pumps, was introduced to allow large scale experiments by allowing a single pressure source to control several models simultaneously. The flow rate controlled by compressed air results in an extremely smooth flow rate with quasi-null flow rate variation”
S.Awwad et al., Pharmaceutics 2023,15, 1444.
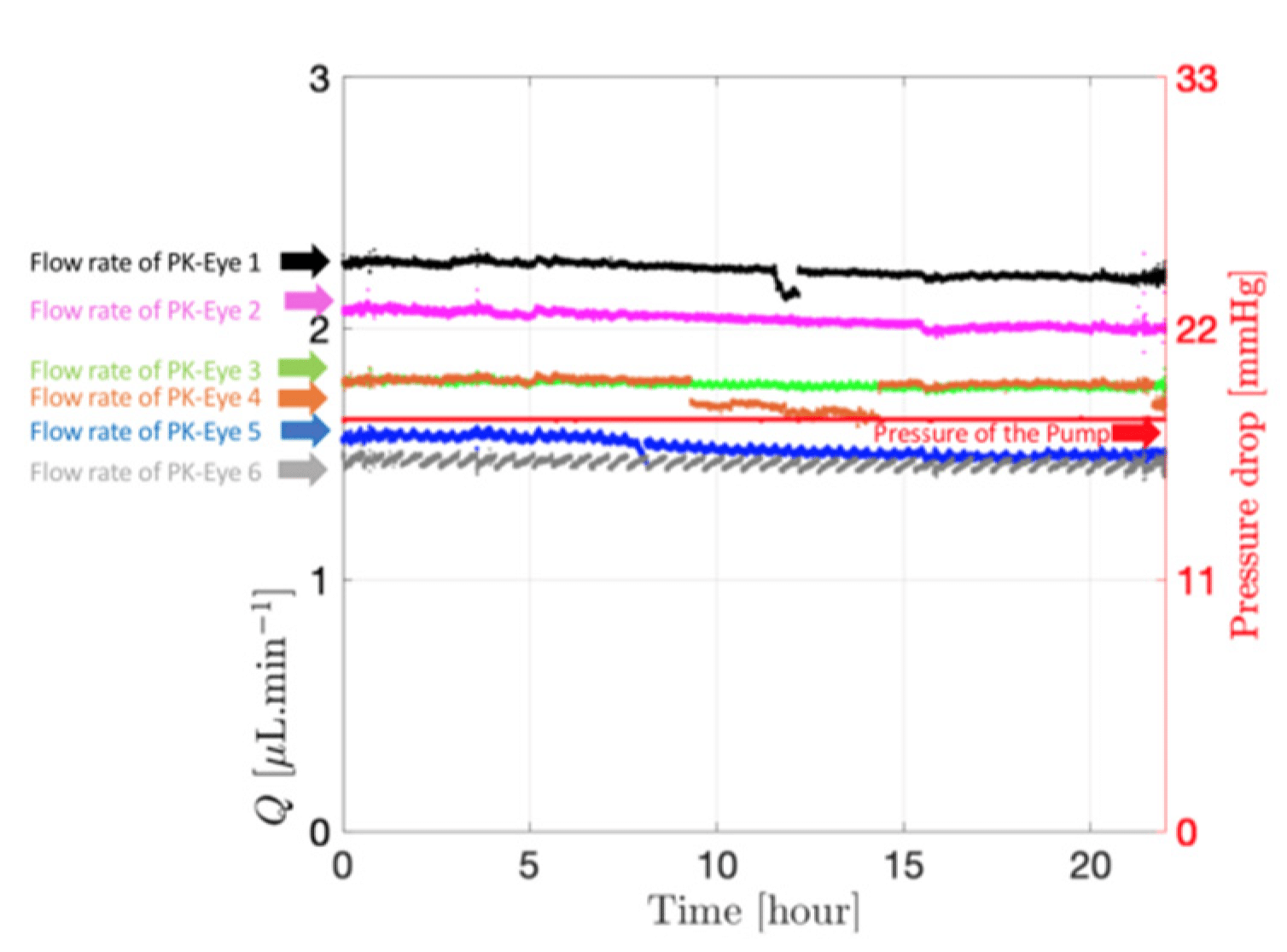
In parallel systems, pressure remained constant, allowing multiple models to be controlled simultaneously from a single pressure source with the ability to run up to six models concurrently. Ratio experiments demonstrated this, with each model flowing within a natural aqueous humor flow rate range (between 1.5 µL/min and 2.7 µL/min), highlighting the uniformity of the setup (figure 7).
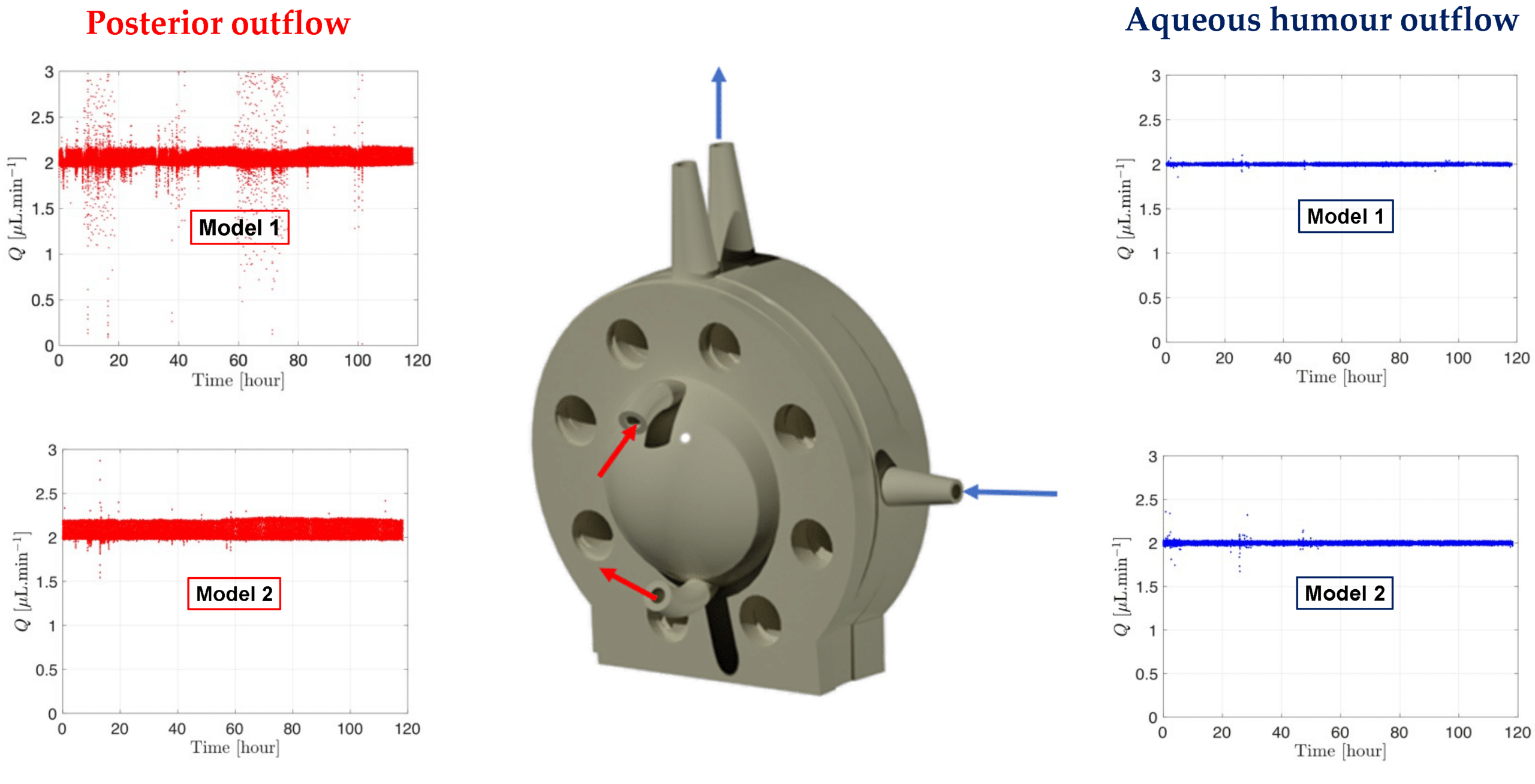

Figure 8: Graph readout from the microfluidic system showing the flow and pressure from the models with SVF at 37°C. [7]
The microfluidic setup with the PK-Eye™ model ensured consistency and detects simulated vitreous fluids (SVF) leakage during testing, critical for maintaining quality control. Stability tests with the model showed consistent pressure and flow rates over 5 days, indicating successful SVF containment without leakage (figure 8). This highlights the robustness of both the microfluidic system and model setup throughout the experiment.
The integration of the 2-SWITCH/M-SWITCH setup facilitated continuous drug quantification across multiple model post-clearance from the PK-Eye™ model, utilizing a connected concentration probe through the M-SWITCH. The platform effectively selected different fluid outlets automatically from this combination of M-SWITCH and 2-SWITCH, enabling real-time monitoring of drug release between multiple flow outlets (figure 9).
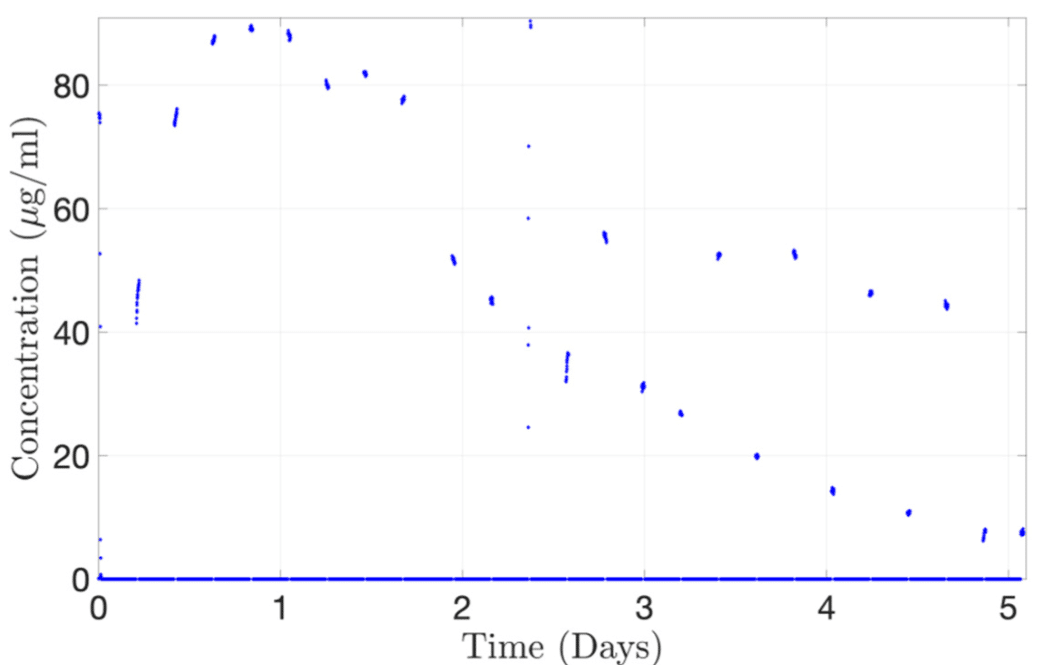
Figure 10: Continuous monitoring of Alexa albumin from the concentration probe connected to 2-SWITCH/M-SWITCH platform.[7]
As a proof-of-concept, the concentration-time profile of Alexa albumin injected into the PK-Eye™ was determined using a concentration probe setup. The platform successfully reproduced the drug release curve, reaching a maximum concentration of approximately 89 μg/mL at 20 hours. This demonstrates the automatic selection of channels flowing through the concentration probe (figure 10). Thus, it highlights the potential for automated redirection of different flows to record the concentration of cleared drugs, showcasing the real-time monitoring capabilities of the platform for ocular drug delivery.
Conclusion
The PK-Eye™ model, developed by Optceutics Ltd., fast-tracks intraocular drug development through real-time monitoring on an automated platform. This platform records pressure, flow, temperature, eye movement, and concentration in connected models, helping in ocular drug delivery studies. Computer-controlled microfluidics mimic ocular flow dynamics with high stability and responsiveness, while an eye movement platform studies their impact on drug clearance. The 2-SWITCH/M-SWITCH platform expands monitoring capacity for multiple models simultaneously.
Related products
Expertise & Resources
-
Microfluidics Article Reviews Microfluidic technology for engineered nanoparticles in nanomedicine Read more
-
Expert Reviews: Basics of Microfluidics Microfluidics for vaccine development Read more
-
Expert Reviews: Basics of Microfluidics Flow control for droplet generation using syringe pumps and pressure-based flow controllers Read more
References:
(1) Ranta, V.; Urtti, A. Transscleral drug delivery to the posterior eye: Prospects of pharmacokinetic modeling. Adv. Drug Deliv. Rev. 2006, 58, 1164–1181.
(2) Jager, R.D.; Aiello, L.P.; Patel, S.C.; Cunningham, E.T. Risks of intravitreous injection: A comprehensive review. Retina 2004, 24, 676–698.
(3) Thrimawithana, T.; Young, S.; Bunt, C. Drug delivery to the posterior segment of the eye. Drug Discov. Today 2011, 16, 270–277.
(4) Souto, E. B.; Dias-Ferreira, J.; López-Machado, A.; Ettcheto, M.; Cano, A.; Camins Espuny, A.; Espina, M.; Garcia, M. L.; Sánchez-López, E. Advanced Formulation Approaches for Ocular Drug Delivery: State-Of-The-Art and Recent Patents. Pharmaceutics 2019, 11 (9), 460.
(5) Laude, A.; Tan, L.E.;Wilson, C.G.; Lascaratos, G.; Elashry, M.; Aslam, T.; Patton, N.; Dhillon, B. Intravitreal therapy for neovascular age-related macular degeneration and inter-individual variations in vitreous pharmacokinetics. Prog. Retin. Eye Res. 2010, 29, 466–475.
(6) Awwad, S.; Henein, C.; Ibeanu, N.; Khaw, P.T.; Brocchini, S. Preclinical challenges for developing long acting intravitreal medicines. Eur. J. Pharm. Biopharm. 2020, 153, 130–149.
(7) Awwad, S.; Ibeanu, N.; Liu, T.; Velentza-Almpani, A.; Chouhan, N.; Vlatakis, S.; Khaw, P. T.; Brocchini, S.; Bouremel, Y. Real-Time Monitoring Platform for Ocular Drug Delivery. Pharmaceutics 2023, 15 (5), 1444.
(8) Liu, T.; Ibeanu, N.; Brocchini, S.; Khaw, P. T.; Bouremel, Y.; Awwad, S. Development of an in Vitro Model to Estimate Mass Transfer from the Anterior Cavity. Front. Drug. Deliv. 2022, 2, 1025029.
A Queen’s University Belfast Paper
Paper: Weaver, E.; Sommonte, F.; Hooker, A.; Denora, N.; Uddin, S.; Lamprou, D. A. Microfluidic Encapsulation of Enzymes and Steroids within Solid Lipid Nanoparticles. Drug Deliv. and Transl. Res. 2023. https://doi.org/10.1007/s13346-023-01398-5.
The Lamprou lab, affiliated with Queen’s University Belfast, specializes in three main areas: nanoparticles for imaging and therapy, lab-on-a-chip technology, and therapeutic implants. Their interdisciplinary approach has driven innovation in healthcare since 2012, developing emerging technologies and novel drug delivery devices. The lab is led by Professor Dimitrios Lamprou, a leading expert in pharmaceutical technologies known for his significant contributions to 3D printing, microfluidics, and nanofibers, with over 150 peer-reviewed publications.The Lamprou lab, in collaboration with the Department of Pharmacy‑Pharmaceutical Sciences (University of Bari Aldo Moro) and Immunocore Ltd., used microfluidics to create eco-friendly lipid-based nanocarriers for encapsulating challenging active pharmaceutical ingredients (APIs).


What are the challenges for biologics and drug delivery?
Biologics are complex molecules derived from living sources, including mRNA vaccines like those used against COVID-19.1,2 Protecting biological drugs from the challenges posed by various inhospitable internal conditions in the body, such as proteases and pH variations, remains a difficult challenge. Since biologics are expensive to produce, optimizing the formulation process is crucial for mass production. There is therefore a growing interest in oral delivery to promote better patient compliance. One promising method is nanoformulation, specifically using solid lipid nanoparticles as a barrier for the active pharmaceutical ingredient post-administration. 3
What makes Solid Lipid Nanoparticles effective for various drug types?
Solid Lipid Nanoparticles have proven their effectiveness in delivering various types of drugs, including chemotherapy drugs, genetic material, and anti-inflammatory medications, through their stability, targeting capabilities, and size.4
SLNs present a special structure with a hydrophobic core and a hydrophilic external layer, enabling them to encapsulate both hydrophilic and hydrophobic substances. The solid core, consisting of solid lipids like waxes and glycerides, is responsible for containing the drugs, particularly hydrophobic ones. On the other hand, the surfactant layer on the outside of an SLN enhances stability and targeted drug delivery and can also contain hydrophilic drugs. The choice of drug significantly influences the components used in lipid-based nanocarrier formulation, with cationic lipid cores and surfactants being important for mRNA and DNA delivery.5
Lipid-based nanoparticles are not limited to SLNs alone, but also include liposomes, niosomes, and exosomes, each with their own distinct properties, as summarized in Table 1.
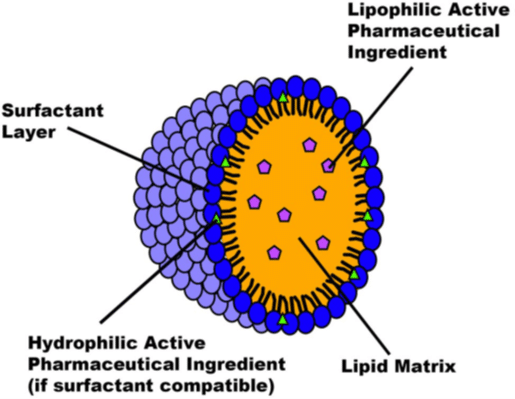
Figure 1: Structure of an SLN capable of encapsulating an API
| Nanoformulation type | Formulation materials | Capable of encapsulating | Advantages | Disadvantages |
| Solid Lipid Nanoparticles | • Waxes • Sterols • Surfactants | • Hydrophobic and hydrophilic | • Highly modifiable • High biocompatibility, including non-toxic degradation | • Require a cooling process for solidification • API leakage during storage • Complications caused by crystallisation |
| Liposomes | • Phospholipids • Cholesterol | • Hydrophobic and hydrophilic | • Highly modifiable • Used for theranostic purposes • Simple synthesis | • Low skin permeability • Infrequently possess low mechanical strength |
| Niosomes | • Non-ionic surfactants • Cholesterol | • Hydrophobic and hydrophilic | • Biocompatible and nonimmunogenic • Improve drug permeation through the skin • Less stringent storage requirements compared to liposomes | • Time-consuming to create • API leakage |
| Exosomes | • Lipids • Proteins • Glycoconjugates | • Hydrophobic and hydrophilic • Genetic material | • Used for theranostic purposes | • Complex and expensive to artificially manufacture |
Table 1: Common lipid-based nanoformulations
How does microfluidics enhance SLN production for drug formulation?
Traditional bulk production methods for solid lipid nanoparticles, such as homogenization and microemulsification, have multiple drawbacks including unpredictable particle characteristics, low reproducibility and repeatability, and the environmentally harmful use of solvents.
To address these issues, microfluidics has emerged as a promising approach for lipid-based nanocarrier formulation and encapsulation of drugs.
Microfluidics has gained popularity in drug delivery due to its precise control of flow rates, device design, and mixing angles within submicron channels. These characteristics enable control over particle size, morphology, and encapsulation efficiency, as flow rates have a significant impact on these parameters. The microfluidic approach also improves time efficiency, allowing for continuous production within minutes, unlike traditional batch processes. As a result, microfluidics is particularly well-suited to formulations relying on self-assembly, like liposomes and SLNs, and has been employed for a wide range of APIs, including those used for gene therapy and addressing the COVID-19 pandemic.6,7,8
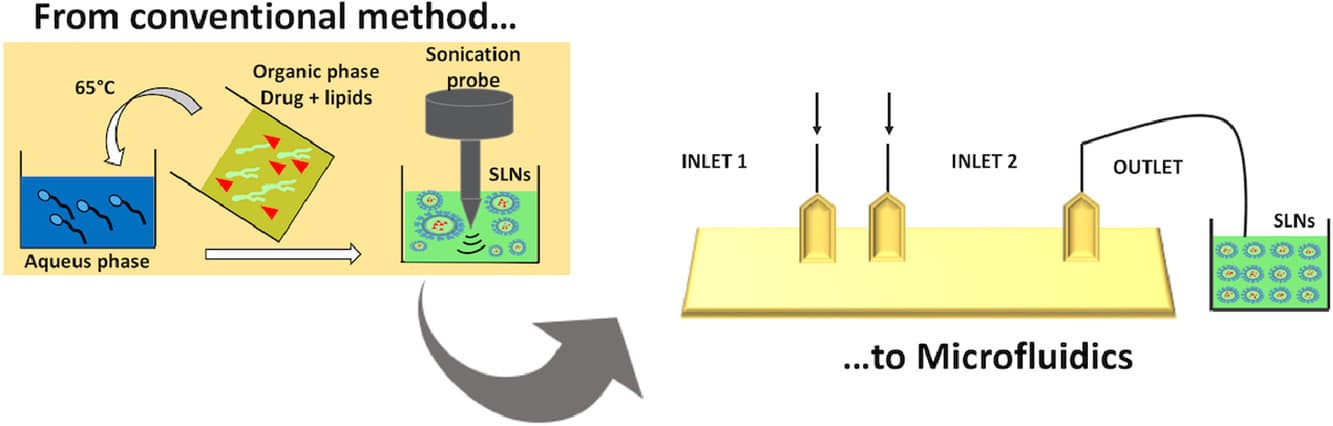
Figure 3 : Microfluidics, a promising approach for SLN nanoformulation.[9]
Aim of the study
To further explore this area of drug encapsulation, Edward Weaver et al. from the Lamprou Lab aimed to demonstrate microfluidics’ position at the forefront of solid lipid nanoparticle production in general, and for biologic SLNs specifically.
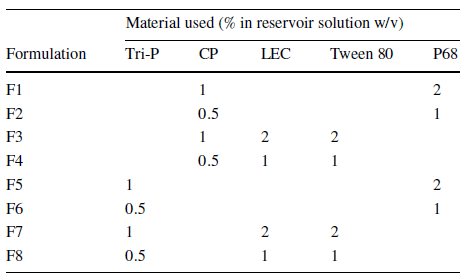
This paper examines the compatibility of trypsin (TRP) and testosterone (TES) for encapsulation in solid lipid nanoparticles using a microfluidic process integrating Fluigent’s Flow EZ. Testosterone was chosen as a positive non-biologic lipophilic API for comparison with previous bulk encapsulation research, while trypsin was evaluated for encapsulating a hydrophilic API in lipid-based nanocarriers. A combination of SLN materials, including tripalmitin (Tri-P), soybean lecithin (LEC), Tween 80 (T80), and cetyl palmitate (CP) in conjunction with Pluronic F68 (P68), was used for nanoformulation using the same experimental setup.
Solid Lipid Nanoparticle Production Method
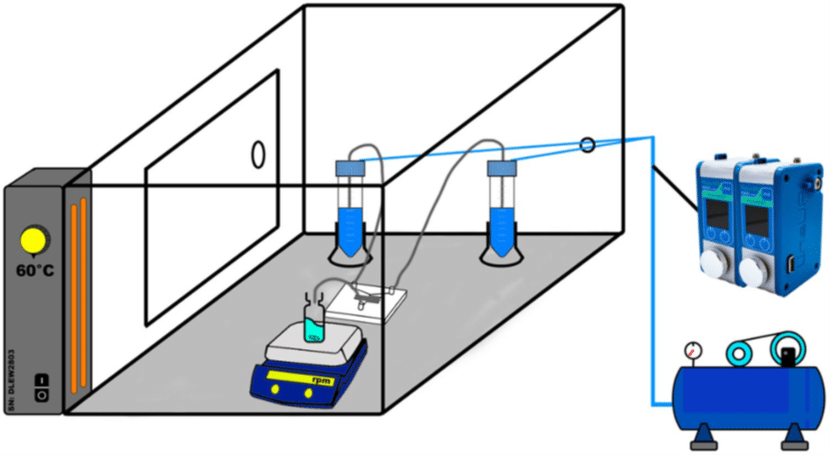
Fluigent’s Flow EZ was used to synthesize various nanoformulations of solid lipid nanoparticles. The microfluidic device used was a Y-shaped inlet with etched herringbone channels. The overall system was maintained at a temperature of 60°C to ensure complete dissolution of the materials.
Eight different SLN formulations (F1-F8) were created, with materials dissolved at 60°C to match their final concentration (Table 2). For SLNs encapsulating APIs, TRP and TES were used at varying concentrations, with TRP in the aqueous phase and TES in the organic phase. The optimal flow rate ratio was determined to be 5:1 (aqueous to organic), and samples were collected. The ethanol excess was evaporated through vortex stirring, and SLN formulations were then refrigerated at 5°C for 24 hours.
The API-loaded SLNs were characterized by dynamic light scattering (DLS), zeta potential, atomic force microscopy (AFM), differential scanning calorimetry (DSC) and Fourier transform infrared spectroscopy (FTIR).
Partial results: Stable and monodisperse SLNs for high encapsulation efficiency.
Solid lipid nanoparticles produced from various combinations and concentrations (F1-F8) were first evaluated by DLS. When comparing non-loaded SLNs to the ones with API (trypsin or testosterone), a slight increase in particle size upon encapsulation was observed. Formulations F2 (CP, P68) and F8 (Tri-P, LEC, Tween 80) presented the most favorable particle sizes (150 -180 nm) and were thus selected as model formulations for the study. The choice of API has a lesser impact on particle size compared to the choice of materials. In fact, the main factor affecting particle size appeared to be the interaction between the waxy core material and the surfactant layer.
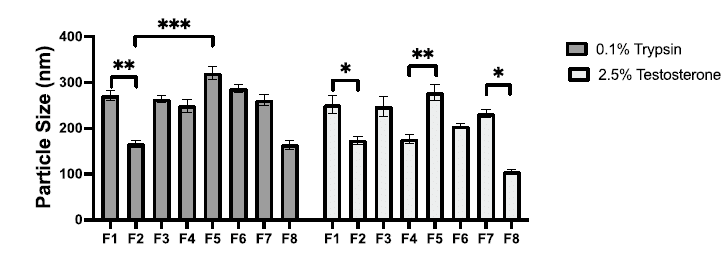
Figure 6 : Particle size measurements for formulations F1–F8.
“However, it was also found in the current study that when using a commercially available chip and the Fluigent system, halving the required concentrations provided more opportune particle diameter. This factor indicates the importance of considering both the system and the microfluidic environment that is being used for formulation. ”
AFM results aligned with DLS analysis, confirming the size range, although slightly enlarged due to drying. It also verified the uniform dispersion of SLNs that was achieved, suggesting great potential for future development.
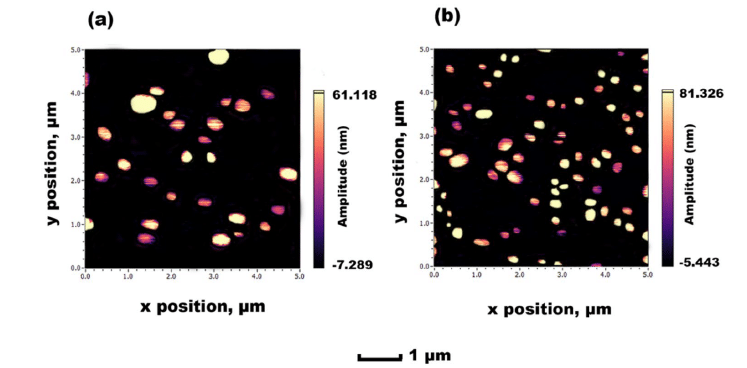
Figure 7: AFM images obtained for F8 SLN encapsulation: (a) TRP and (b) TES
Regarding stability studies, all of the formulations (F1-F8) appeared to function effectively. In particular, F2 and F8 demonstrated favorable stability characteristics, ensuring consistent and prolonged release.
FTIR spectroscopy confirmed the presence of TRP and TES in SLN formulations. Specific peaks in the spectra indicated the presence of these compounds in the formulations, thus demonstrating their encapsulation within the solid lipid nanoparticles.
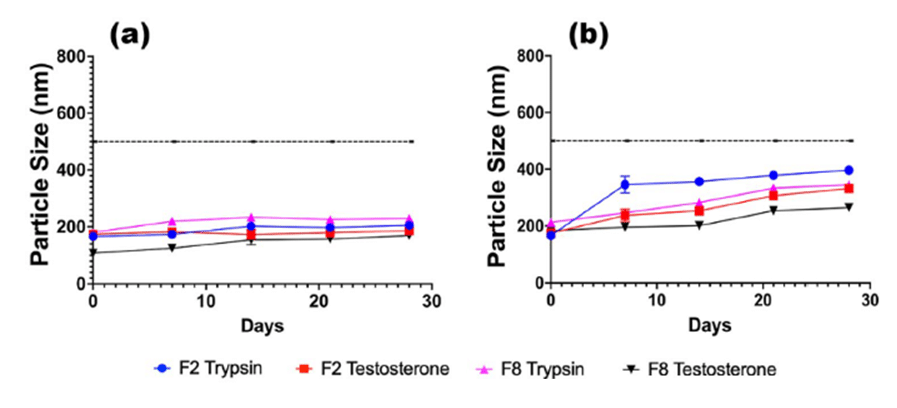
Figure 8: Stability for TRP and TES-loaded F2 and F8 formulations, stored at a) 5 °C and b) 37 °C
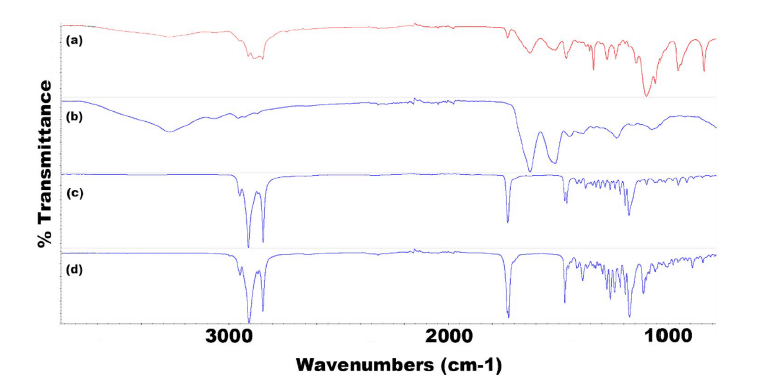
Figure 9: FTIR spectra for all components of F2 TRP showing (a) F2 TRP-encapsulated SLNs, (b) TRP, (c) CP, and (d) P68
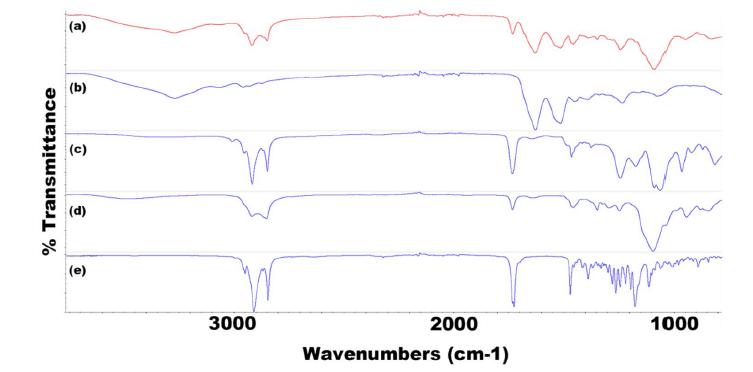
Figure 10: FTIR spectra for all components of F8 TRP showing (a) F8 TRP-encapsulated SLNs, (b) TRP, (c) LEC, (d) T80, and (e) Tri-P
Finally, the encapsulation efficiency (EE) and drug release with F2 and F8 formulation were evaluated. For testosterone, the encapsulation efficiency was high for both nanoformulations, showing a slight improvement over traditionally used encapsulation methods such as emulsification and homogenisation.10,11 Moreover, the release of TES from SLNs was around 65% for F2 and 45% for F8 within 72 hours, in agreement with the therapeutic range.
Regarding the encapsulation of TRP within solid lipid nanoparticles, the EE with the F2 formulation was 47% and showed controlled trypsin release, making it a suitable lead formulation compared to F8 (EE: 7%). This result is promising for future enhancement, taking into consideration that this encapsulation had not been attempted before with microfluidics.
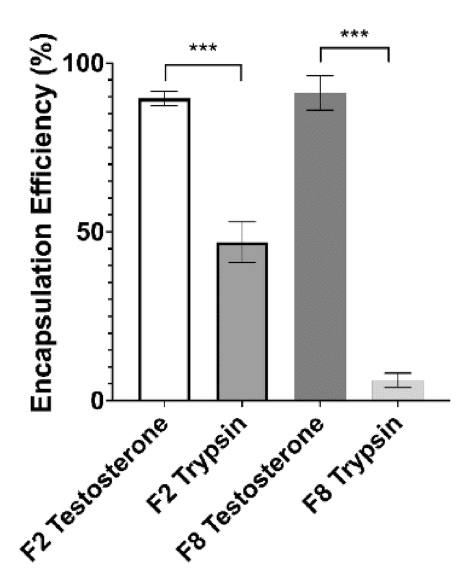
Figure 11: Encapsulation efficiency for F2 and F8 formulations encapsulating both TRP and TES
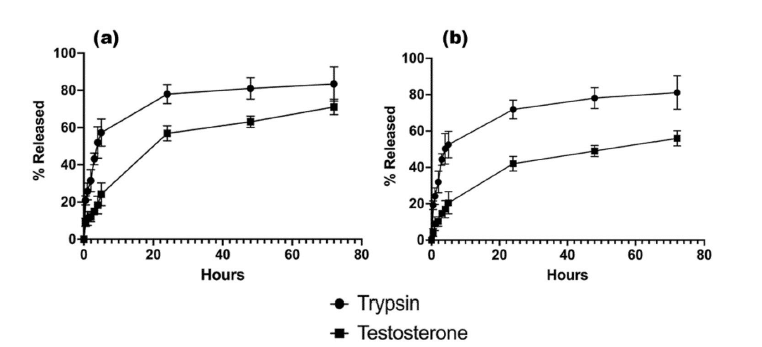
Figure 12: Drug release displayed as % total release from encapsulated active pharmaceutical ingredient for (a) F2 and (b) F8
Conclusion
In this paper highlight, E. Weaver et al. from the Lamprou lab developed a microfluidic approach for solid lipid nanoparticle production and API encapsulation. Fluigent’s Flow EZ was used to provide controlled and constant flow rates, contributing to the creation of small homogeneous SLNs. Selecting appropriate SLN materials was essential to matching specific APIs due to their varying encapsulation capabilities. The nanoformulation which included low-concentration CP and P68 was shown to be the most promising.12 This microfluidic method was both reproducible and eco-friendly, making it possible to encapsulate previously challenging molecules.
Related products
Related Content

Microfluidics for Pharmaceutical Applications
Read more
Webinar – Microfluidics in the manufacturing of sustainable nanomedicines
Read more
Expertises & Resources
-
Expert Reviews: Basics of Microfluidics Microfluidics in Drug Delivery: A New Era of Precision Medicine Read more
-
Microfluidic Application Notes 1-10 microns PLGA microsphere production using the RayDrop Read more
-
Microfluidics Article Reviews A mRNA encapsulation platform integrating Fluigent’s FlowEZ Read more
-
Microfluidics Article Reviews Microfluidic technology for engineered nanoparticles in nanomedicine Read more
-
Microfluidic Application Notes PLGA microcapsules synthesis Read more
-
Expert Reviews: Basics of Microfluidics Microfluidics for vaccine development Read more
-
Microfluidic Application Notes Liposome Nanoparticle Synthesis Read more
-
Expert Reviews: Basics of Microfluidics Flow control for droplet generation using syringe pumps and pressure-based flow controllers Read more
References
- (1) Anselmo AC, Gokarn Y, Mitragotri S. Non-invasive delivery strategies for biologics. Nat Rev Drug Discovery. 2019;18(1):19–40.
- (2) Schoenmaker L, et al. mRNA-lipid nanoparticle COVID-19 vaccines: structure and stability. Int Pharm. 2021;601: 120586 .
- (3) Mishra V, et al. Solid lipid nanoparticles: emerging colloidal nano drug delivery systems. Pharmaceutics. 2018;10(4):191.
- (4) Mura P, et al. Evaluation and comparison of solid lipid nanoparticles (SLNs) and nanostructured lipid carriers (NLCs) as vectors to develop hydrochlorothiazide effective and safe pediatric oral liquid formulations. Pharmaceutics. 2021;13(4):437.
- (5) Sommonte F, et al. The complexity of the blood-brain barrier and the concept of age-related brain targeting: challenges and potential of novel solid lipid-based formulations. J Pharm Sci. 2022;111(3):577–92.
- (6) Jaradat E, et al. Microfluidic paclitaxel-loaded lipid nanoparticle formulations for chemotherapy. Int J Pharm. 2022;628:122320.
- (7) Jain V, et al. Microfluidic device based molecular self-assembly structures. J Mol Liq. 2022: 119760.
- (8) Roces CB, et al. Manufacturing considerations for the development of lipid nanoparticles using microfluidics. Pharmaceutics. 2020;12(11):1095
- (9) Arduino I, et al. Preparation of cetyl palmitate-based PEGylated solid lipid nanoparticles by microfluidic technique. Acta Biomater. 2021;121:566–78.
- (10) Tajbakhsh M, et al. An investigation on parameters affecting the optimization of testosterone enanthate loaded solid nanoparticles for enhanced transdermal delivery. Colloids Surf A. 2020;589:124437.
- (11) Doktorovova S, Souto EB, Silva AM. Hansen solubility parameters (HSP) for prescreening formulation of solid lipid nanoparticles (SLN): in vitro testing of curcumin-loaded SLN in MCF-7 and BT-474 cell lines. Pharm Dev Technol. 2018;23(1):96–105.
- (12) Weaver, E.; Sommonte, F.; Hooker, A.; Denora, N.; Uddin, S.; Lamprou, D. A. Microfluidic Encapsulation of Enzymes and Steroids within Solid Lipid Nanoparticles. Drug Deliv. and Transl. Res. 2023.
Benefit from the best performance and Fluigent’s expertise for your project
- High performance: pressure-based flow control allows for highly responsive and stable pulseless flow
- Versatility: perform any protocol including injection, uni-directional recirculation, sampling, or perfusion
- Transportability: compactly packaged, the pressure source, pressure controllers, flow rate sensors, and fluidic paths – the entire unit can be customized with injection molding techniques
- Sterility: does not require users to manipulate the chip or the system
Microfluidic recirculation system: technology description
Pressure-based flow control for stable pulseless recirculation
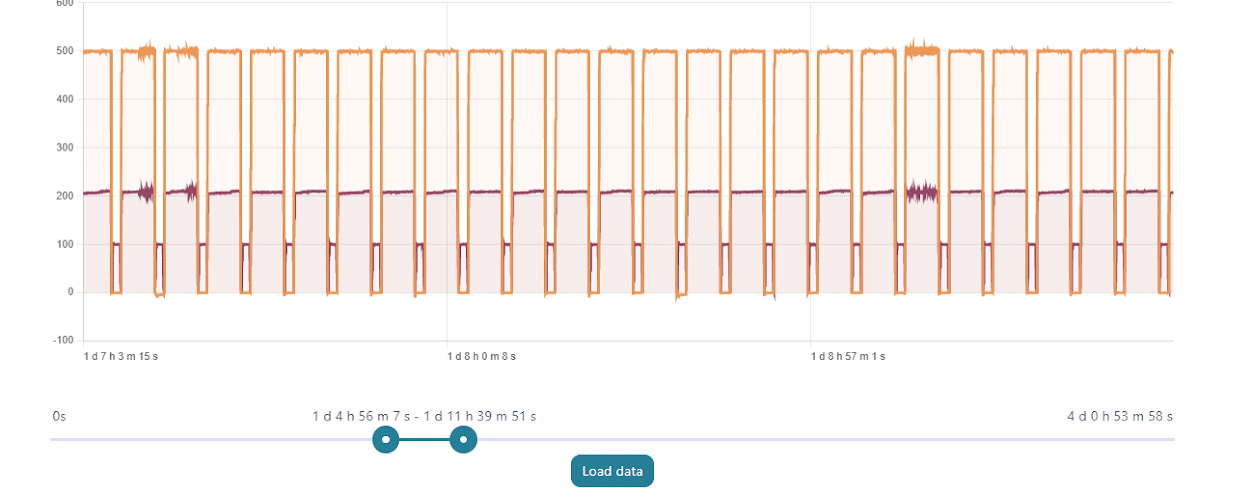
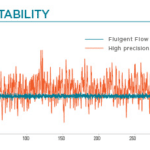
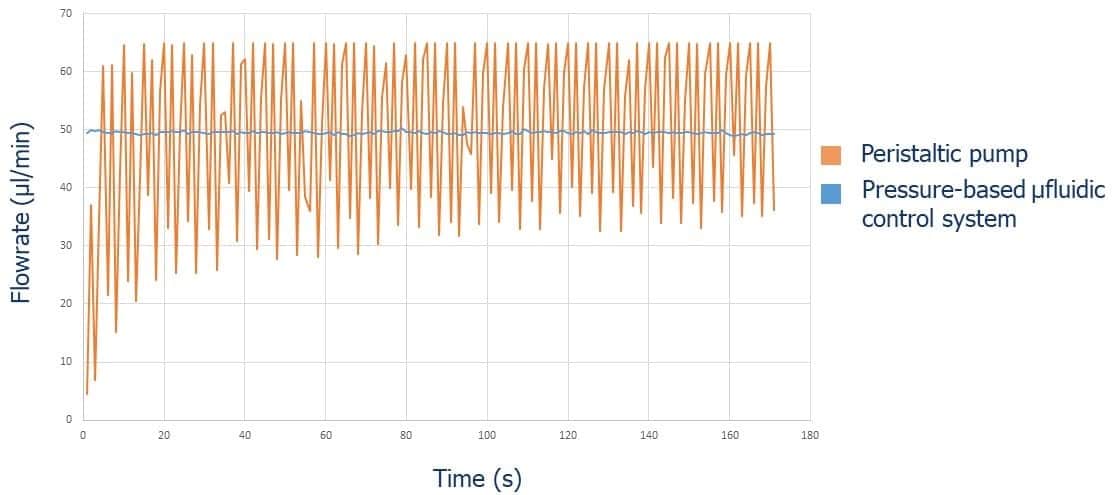
Recirculation may be accomplished with a peristaltic pump or pressure controller. The latter has the advantage of providing a more accurate and stable flow profile. The flow from the peristaltic pump leads to pulsatile flow rates with oscillations of more than 40% of the set value (50 µL/min, see graph below). Fluigent pressure controllers have a negligible flow variation with improved response time. Using a peristaltic pump can increase the risk of cell detachment, loss of membrane integrity, and/or cell dysfunction due to pulsation. Pressure-based flow controllers offer minimal shear stress comparable to that in an in vivo environment.
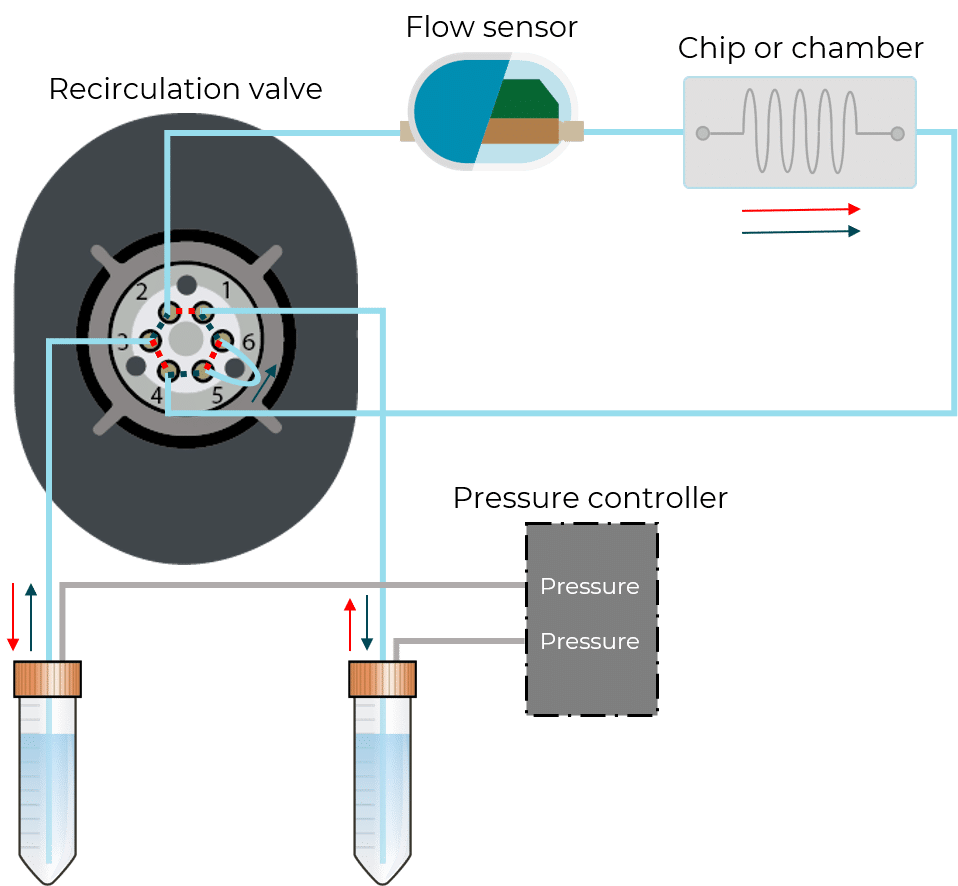
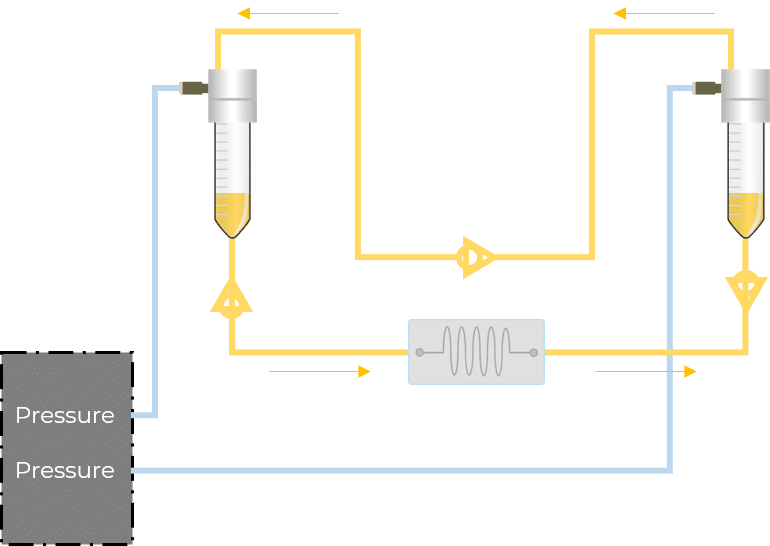
For microfluidic recirculation using pressure-based flow control, one requires a pressure source, a pressure controller with 2 channels, a 2-position 6-port valve, and a flow rate sensor. This allows for flow between two reservoirs while maintaining a continuous unidirectional flow rate in the chamber.
Although this solution is functional and offers highly improved performance, additional efforts should be made in order to mitigate the limitations of using such systems compared to a peristaltic pump, including the additional liquid volume required for this setup, the complexity and footprint, and the overall system cost.
Fluigent’s new technology allows for automatic refills so that perfusion can be performed continuously
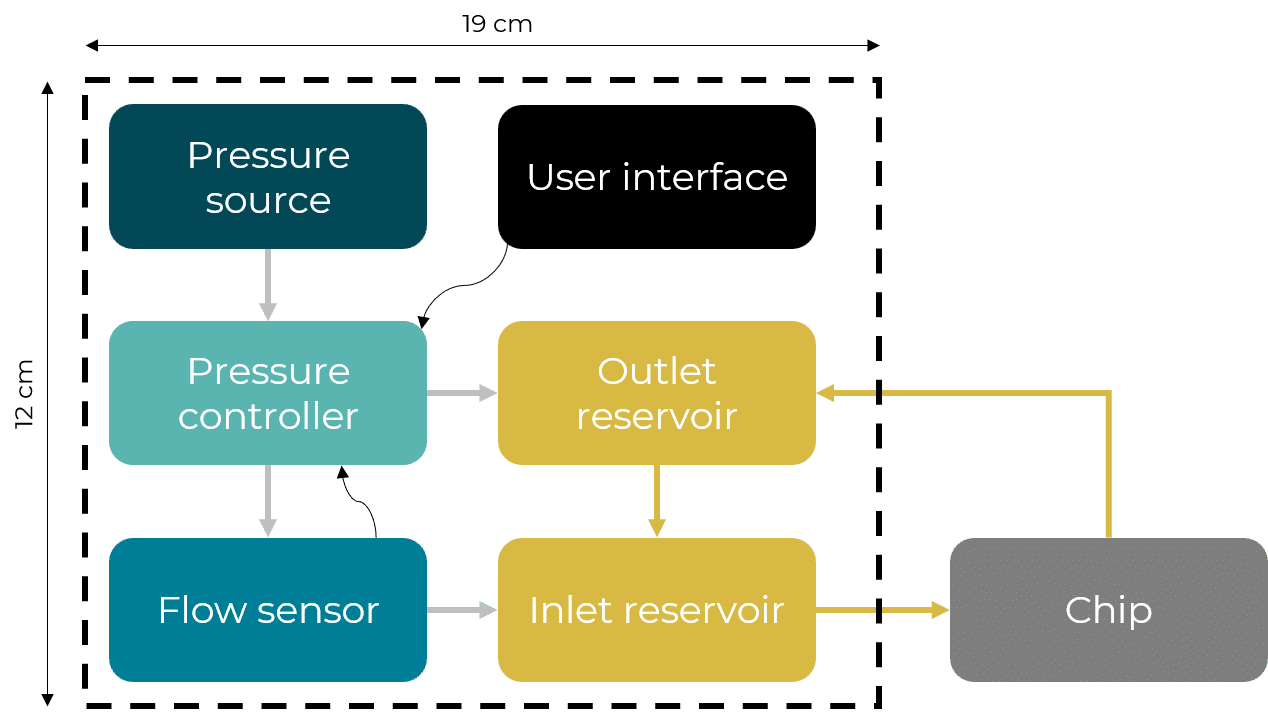
The microfluidic recirculation system directly monitors the volume left in the reservoirs and integrates all components needed in a functional microfluidic recirculation system in one compact device (that can be used with a microscope or in an incubator): pressure source, pressure-based flow controller, flow sensor (for flow rate regulation), reservoirs, all the tubing and electrical connections necessary, and a user interface (touchscreen). Users only have to provide the microfluidic module with the needed chip for their application.
Figure 2: Recirculation classical setup vs Fluigent all-integrated system
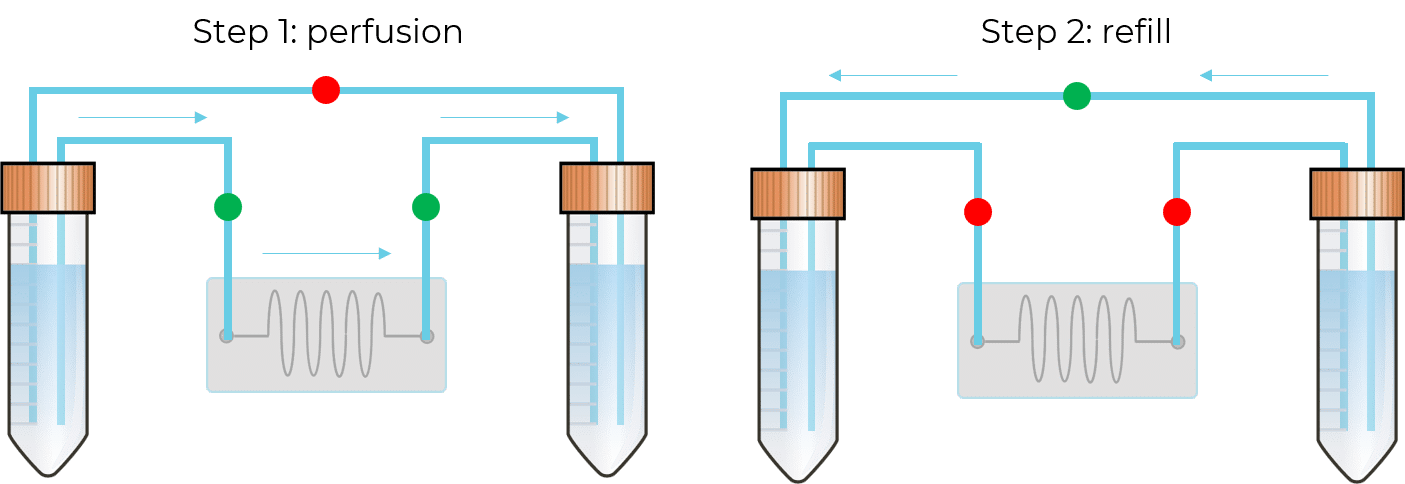
In this configuration, there is no need for a 2-position valve. Instead, there are two fluidic paths: one that goes through the chip or chamber, and one that directly links the reservoirs. In each path, the flow is feeding through in one direction with the use of valves. These enable the system to have flow going from one reservoir to the other only when required.
Optical detection for long-term flow rate reliability
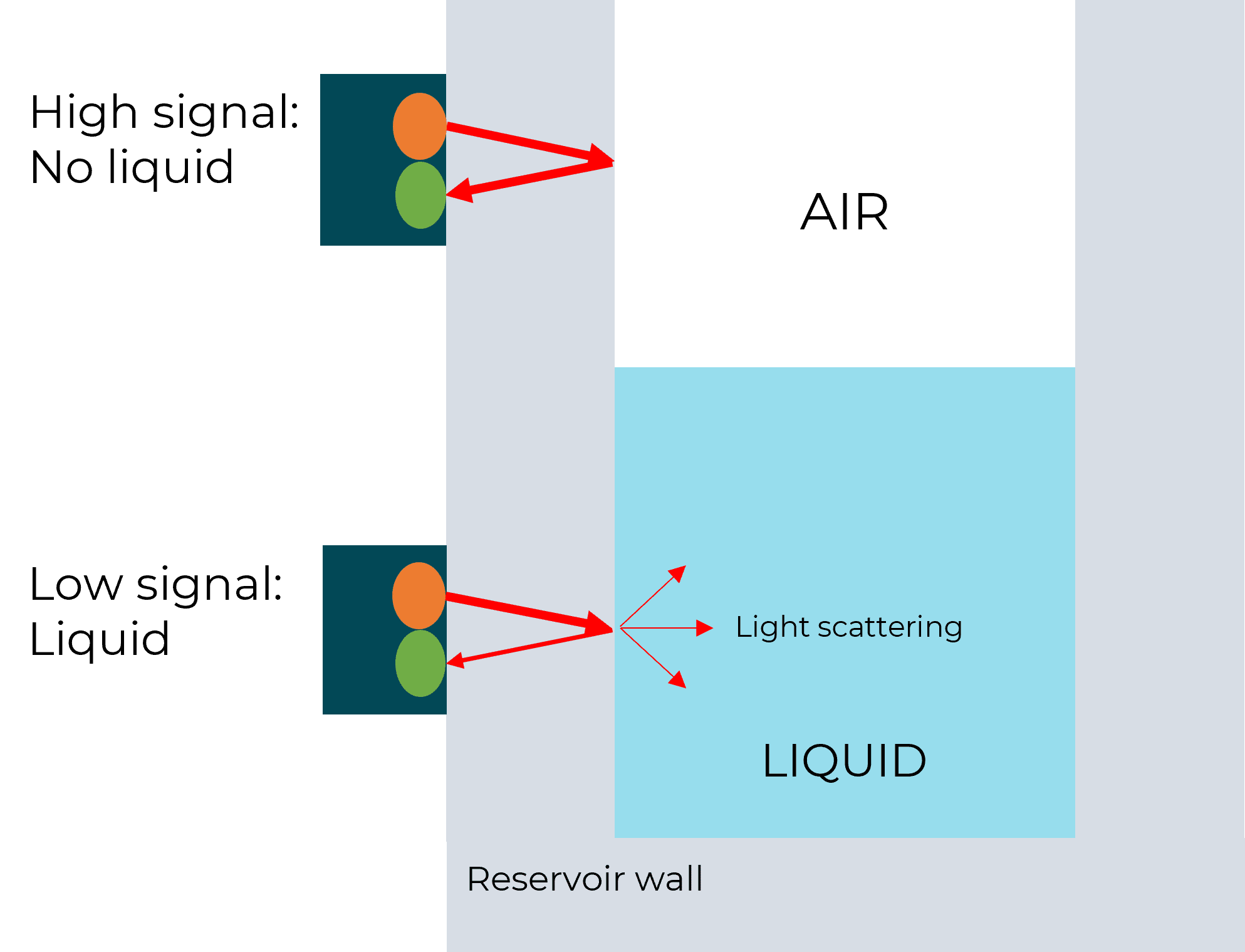
To automate the recirculation process, the device requires the ability to know the liquid level when the inlet reservoir is almost empty, or the outlet reservoir is almost full. The solution, developed by Fluigent, is based on optical level sensors (patented), it detects the presence or the absence of liquid in the reservoir.
One, or several sensors can be mounted at different heights to detect the level of liquid for which the reservoir is considered “in need of refill”. Fluigent algorithms then adjust the pressure applied to the outlet recirculation reservoir to reach the check valve’s threshold and ensure a fast refill from the outlet reservoir to the inlet reservoir while interrupting the pressure applied to the inlet reservoir for this short while.
The liquid volumes measured by the optical sensors allow users to ensure the correct flow rates are measured with the flow sensors. The volume of solution in the reservoir, according to the measured flow rate and elapsed time, is compared to the information given by the flow sensor. If a drift is detected, the flow sensor will be automatically recalibrated.
Unpluggable cartridge
Fluigent developed disposable cartridges to be used in microfluidic recirculation systems. They allow sterility and reproducibility for biological applications that require a contamination-free environment. Fluigent can also provide disposable and micro-patterned polymer-based fluidic parts that can be packaged and sterilized for medical applications.
The fabrication technology consists of a multiplayer bonding of plastic parts that can be either molded or machined to include valves to prevent backflow in the recirculation path (patented).
These components are easily integrated into a larger module with a one-click interconnection and locking technology (patented). Through an ingenious system,the cartridge can be placed and locked in its position or unlocked and removed easily. When the cartridge is in place, it is leakly-tight connected to the base manifold. Thus, neither air nor liquid can flow throughout the gasket.

Automation and software integration
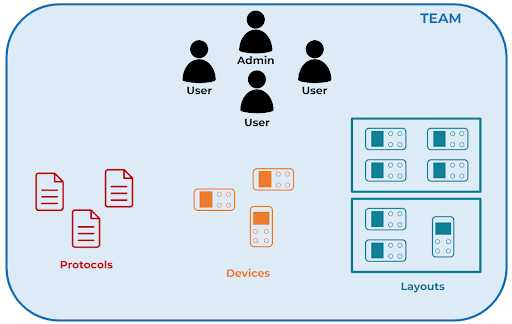
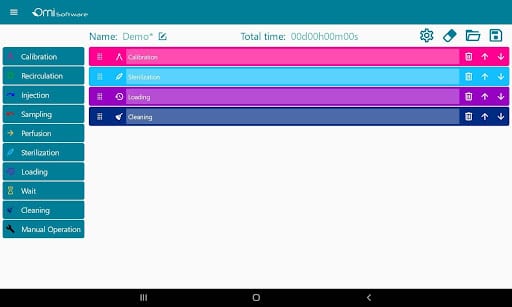
The software can manage several recirculation modules from individual users and teams.
Users can directly write protocols via the interface as it is equipped with predefined steps such as injection, perfusion, sampling, or recirculation that help users build protocols. These steps are editable, and users have the ability to choose parameters such as volumes, flow rates, durations, or flow patterns.
The data can be visualized and analyzed via graphs in real time and afterward.
Fluigent developed a solution for remote control of the microfluidic recirculation system using WI-FI connectivity. It is possible for us to design a custom Android app to edit protocols, pair, and register devices, monitor experiments, and visualize results. These functionalities can also be implemented in a web interface so that it is reachable from any device that supports a browser.
For those who prefer on-site control or want more control, Fluigent can make an embedded touchscreen included in the microfluidic recirculation system with a dedicated interface with access to basic controls.
As for data management, Fluigent provides a cloud service to collect and store data that can be retrieved when needed.
Performance of Fluigent microfluidic recirculation system
With a set perfusion flow rate of 500 µL/min, the recirculation module can maintain a constant flow rate with regular refill steps for days without any offset or issue related to the emptying of the reservoirs. On the graph above, one can see the evolution of the flow rate during 7-hour portions of a 4-day experiment.
Microfluidic applications related to microfluidic recirculation system
Dynamic cell culture
Long-term culture of cells requires the continuous perfusion of medium and thus recirculation in the fluidic path. It is interesting to have an automated and maintenance-free solution for days-long or weeks-long processes. This process must be efficient to guarantee the conservation of cells in an environment with the right amount of reagents for the entire duration of the manipulation. [1][2][3]
Live cell imaging and bioanalysis
To optically analyze live cells, scientists must keep the cells in given physiological conditions during the analysis while minimizing manual manipulations required for reagent injection, media exchange, or sample collection. Therefore, they need a module that is compact enough to be compatible with screening devices that also integrates a cell media microfluidic recirculation system to maintain the cells in the desired state over the longer term. [3][4][5]
Organ-on-a-chip
The combination of a microfluidic recirculation system with microfluidic devices can model liquid/liquid or liquid/air interfaces and mimic physiological behaviors of different human organs such as gut, lung, or heart cells inside a microchip, which is useful for drug discovery and diagnostics.
- For organ-on-a-chip-based drug discovery, recirculating media allows continuous perfusion that reproduces natural phenomena in the human body. When looking for treatment against diseases, the technology is useful to study the organs’ response to drugs in the preclinical stage to identify and validate them and prevent clinical failures later on. [6][7]
- Organ-on-a-chip development also paves the way for applications related to immunoassays, nucleic acid, or cell testing with automation, parallelization, and miniaturization of the processes. This is beneficial for point-of-care diagnostics, as the use of microfluidic recirculation permits rapid biomarker detection and thus, diagnosis of infectious diseases from sera extracted from blood with reduced costs. [8][9]
Related products

Omi, an Automated Organ-On-A-Chip Platform
Mimic Microphysiological Conditions in Organ-on-a-Chip Studies
See the offer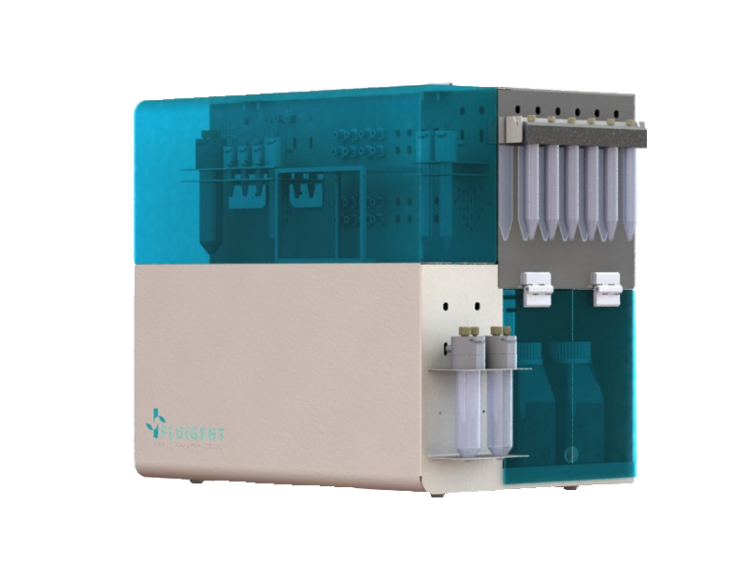
Fully Custom Microfluidic Device
Fully Custom Microfluidic Device
See the offer
Microfluidic OEM Pressure Controller
OEM Fluigent PX
See the offer
Microfluidic OEM Flow Sensor
FS Series
See the offer
Ressources
References
[1] A. R. Dixon, S. Rajan, C.-H. Kuo, T. Bersano, R. Wold, N. Futai, S. Takayama, G. Mehta, “Microfluidic device capable of medium recirculation for non-adherent cell culture”, Biomicrofluidics, 2014
[2] K. B. Chang, A.-S. Kameel, Y.-K. Cho, T. Shuichi, “Pumps for microfluidic cell culture”, Electrophoresis, 2014
[3] N. Futai, W. Gu, J. W. Song, S. Takayama, “Handheld recirculation system and customized media for microfluidic cell culture”, Lab Chip, 2006
[4] C. M. Puleo, H. C. Yeh, K. J. Liu, T. H. Wang, “Coupling confocal fluorescence detection and recirculating microfluidic control for single particle analysis in discrete nanoliter volumes”, Lab Chip, 2008
[5] F. Cantoni, G. Werr, L. Barbe, A. M. Porras, M. Tenje, “A microfluidic chip carrier including temperature control and perfusion system for long-term cell imaging”, HarwareX, 2021
[6] D. R. Taft, “The Isolated Perfused Rat Kidney Model: A Useful Tool for Drug Discovery and Development”, Current Drug Discovery Technologies, 2004
[7] E. W. Esch, A. Bahinski, D. Huh, “Organs-on-chips at the frontier of drug discovery”, Nature Reviews Drug Discovery, 2015
[8] J. Ducrée, “Next-generation microfluidic lab-on-a-chip platforms for point-of-care diagnostics and systems biology”, Procedia Chemistry, 2009
[9] N. Garg, D. Vallejo, D. Boyle, I. Nanayakkara, A. Teng, J. Pablo, X. Liang, D. Camerini, A. P. Lee, P. Felgner, “Integrated On-Chip Microfluidic Immunoassay for Rapid Biomarker Detection”, Procedia Engineering, 2016
Table of contents
I. Droplet-based microfluidics, a microfluidic chip application
Microfluidic generation of droplets has attracted a lot of interest, enabling high throughput experiments by generating millions of micro-reactors and chambers in a few seconds. This microfluidic chip application method produces highly monodispersed droplets of very small volumes (μL to fL) of fluids with high frequency (up to hundreds of kHz), providing better control of processes like mixing, encapsulation, sorting, and sensing. Microfluidic-based droplets have many diverse and varied applications, such as particle synthesis3 and chemical analysis4. Highly controlled droplet production also enables single-cell analysis or drug testing 5,6.
Microfluidic-based droplet generation and control allow for:
- Highly monodispersed (<2% size variation) droplet production, with potentially high generation rate (up to hundreds of kHz)
- Highly reproducible complex structures (water-in-oil-in-water emulsions, multiple encapsulations…)
- Single droplet manipulation as an individual pL scale biochemical reactor
- Miniaturization of production and bioanalytical devices
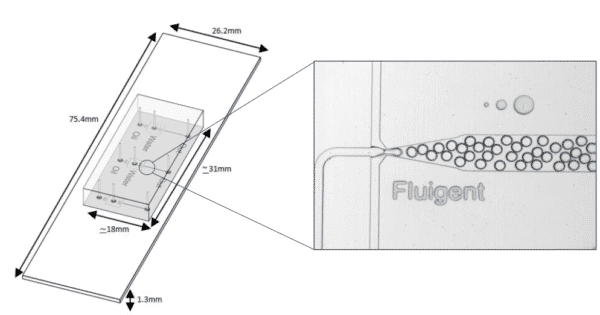
Figure 1: The Fluigent EZ Drop for droplet generation
With these characteristics, droplet microfluidics has a large value, including bio(chemical) analysis, and nano and microscale generation of materials7. Several microfluidic chip designs exist to generate droplets for a various field of applications. A common design is the T-junction, where the dispersed phase is injected from a channel that is perpendicular to the channel carrying the continuous phase (figure 1).
Application of microfluidic chip using droplet microfluidic concepts, components, and processes are now being adopted and leveraged by end-users to enable new science and innovation. Real-world success is now evidenced through a range of mainstream commercial products that are applied to key biological and healthcare-related problems (e.g., 10X Genomics, Drop-seq, and nucleic acid quantification via Droplet DigitalTM PCR systems)8.
Today, it is possible to produce multiple emulsions with complex droplet morphologies. Production of multi-cored droplets (droplets that contain a controlled number of inner droplets at one or more hierarchical levels), Janus droplets (i.e. biphasic or triphasic droplets with two or three physically and chemically distinct surface domains) is now possible using droplet microfluidics9 (figure 2).
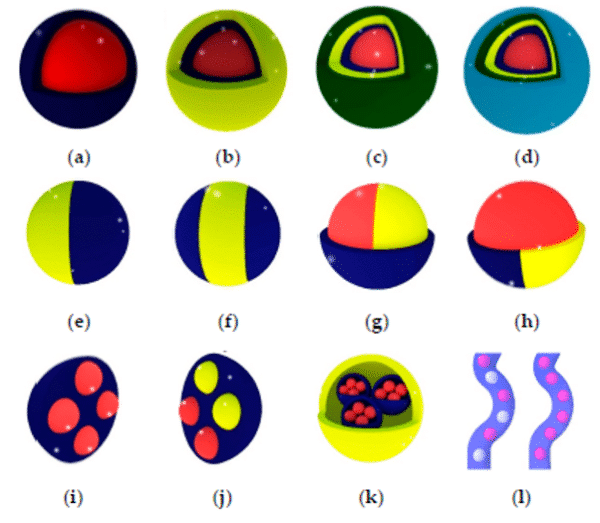
Figure 2: Types of multiple emulsions from the simple encapsulation (a), double emulsion (b), and so on (c,d) to bi- and tri-phasic structures (e to h), multiple encapsulations (I to k), and hybrid microjet achievable using droplet-based microfluidics9
Microfluidic chip application for the production of highly reproducible PLGA microparticles
In recent years, biodegradable microspheres/microparticles have gained widespread importance in the delivery of bioactive agents10. The copolymers of Poly (D, L-lactic-co-glycolic acid) (called PLGA)/poly (lactic acid) PLA microparticles are one of the most successful new drug delivery systems (DDS) in labs and clinics. Because of their biocompatibility and biodegradability, they can be used in various areas, such as long-term release systems, vaccine adjuvant, and tissue engineering11.
In PLGA microparticle production for drug release and delivery, microparticle size is a key parameter as it is directly related to the microparticles degradation rate and the drug release rate12. Although PLGA microparticle synthesis appears to be a successful drug delivery system, the current processes and tools to produce PLGA microparticles have many limitations, such as wide microparticle size distribution, poor repeatability, and aggressive chemical preparation conditions11. To solve these problems, droplet-based microfluidics application chip offers an efficient method for improvement.
Fluigent provides the Raydrop: a new breakthrough technology leading to outstanding particle size monodispersity and production flexibility. The Raydrop works as a co-flow focusing principle (figure 3). The nozzle and outlet capillary are aligned in a continuous phase chamber, the dispersed phase comes through the nozzle to create the microparticles into the continuous phase and exits by the outlet insert. Using this application of microfluidic chip method, PLGA microparticles ranging from 15 to 50 µm diameters have been successfully generated.
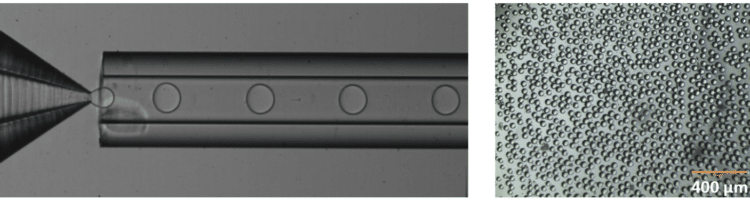

The PLGA microparticle production station allows excellent reproducibility and significantly improved monodispersity (CV < 2%) as compared to other methods on the market. It allows one to continuously produce PLGA microparticles (up to 10 000 droplets/s) without unwanted interruption for long-term experiments.
Apllication notes:
A microfluidic chip application for single-cell mRNA-seq sequencing using droplets: Drop-seq technology
The production of highly monodispersed emulsions or more complex structures (water-in-oil-in-water emulsions, multiple encapsulations…) makes microfluidic chip applications an excellent approach for single-cell analysis or single-cell culture. The technique allows for droplet-based single-cell RNA-sequencing, such that one can characterize complex tissues with many cell types and states under diverse conditions. One of the pioneering microfluidic chip application methods is Drop-Seq technology, which entraps a single cell and a single primer-barcoded bead in each droplet (figure 4).
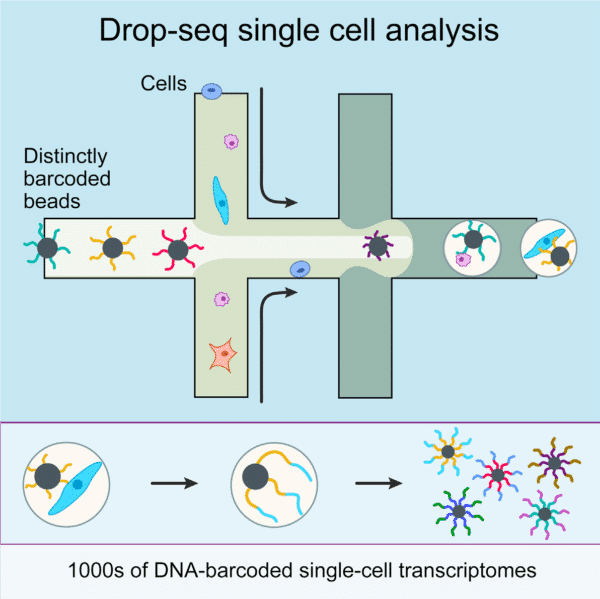
Figure 4: Scheme representing Drop-Seq single cell analysis. A microliter droplet encapsulates a cell and a barcoded bead for identification13
Cells are thus separated into nanoliter-sized aqueous droplets, with a different barcode associated with each cell’s mRNAs. The primers on beads contain a barcode consisting of three sequences. One sequence is for PCR amplification and is common to all the beads. The second sequence consists of hundreds of individual primers that also share the same ‘‘cell barcode’’.
Finally, the third part has different unique molecular identifiers (UMI), enabling mRNA transcripts to be digitally counted13 (figure 5). The droplets are sequenced altogether, allowing quick profiling of thousands of individual cells from a heterogeneous population.
The power of this microfluidic chip application technology resides in the fact that during sequencing, one can distinguish where the original information came on a cell to cell basis.
This allows one to make a gene expression map of the cell, or even to distinguish cell populations within a tissue.
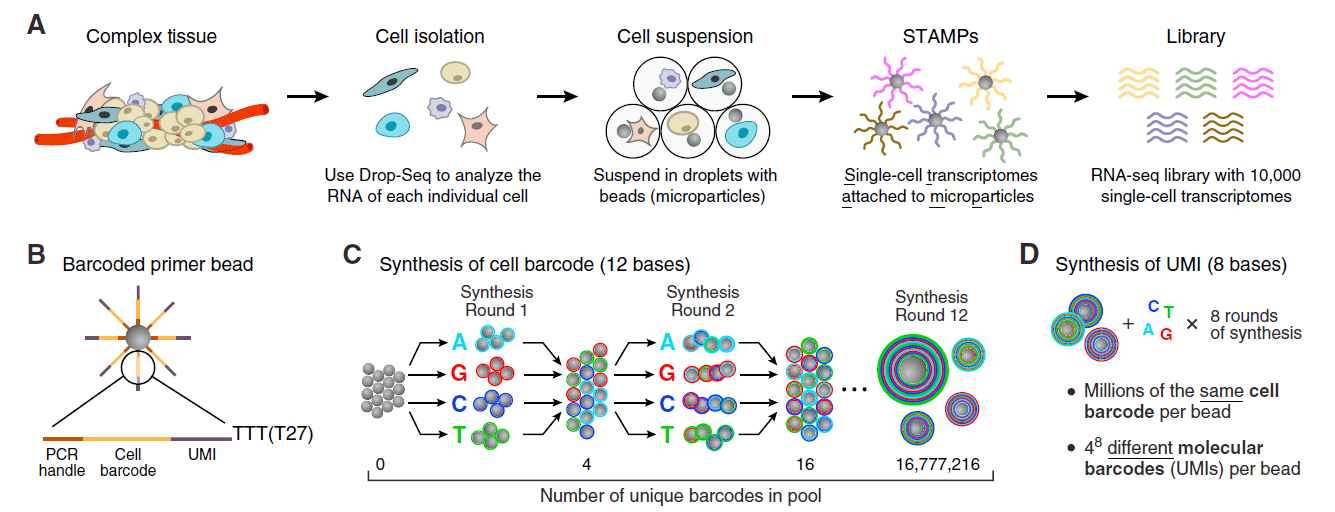
Figure 5: Molecular barcoding of cellular transcriptomes in droplets microfluidics: (A) Drop-Seq barcoding schematic, (B) Sequence of primers on the microparticle, (C) Split-and-pool synthesis of the cell barcode, and (D) Synthesis of a unique molecular identifier13
Apllication notes:
II. Microfluidic cell culture for a better cell behavior understanding
Microfluidic cell culture is another application of microfluidic chip that has significant advantages over macroscopic culture in flasks, dishes, and well-plates14 (figure 6). The microfluidic chip fabrication process allows great flexibility in the design of microfluidic devices, permitting one to understand and control interactions between cells, substrates, and the surrounding medium, physically as well as biochemically15.
This microfluidic chip application technology offers new possibilities to accurately reproduce the cellular environment and enables the analysis of biological processes that were not accessible before. Morphology-wise, chips can be structured at the cell scale to reproduce the mechanical constraints experienced by cells. Biochemically, stable gradients can be implemented with a high spatial resolution (typically, micrometer resolution).
Finally, constant perfusion enables the continuous renewal of nutrients and oxygen to promote cell growth and maintain optimal activity during long-term cell culture. Cost reduction due to volume reduction is also a major benefit15.
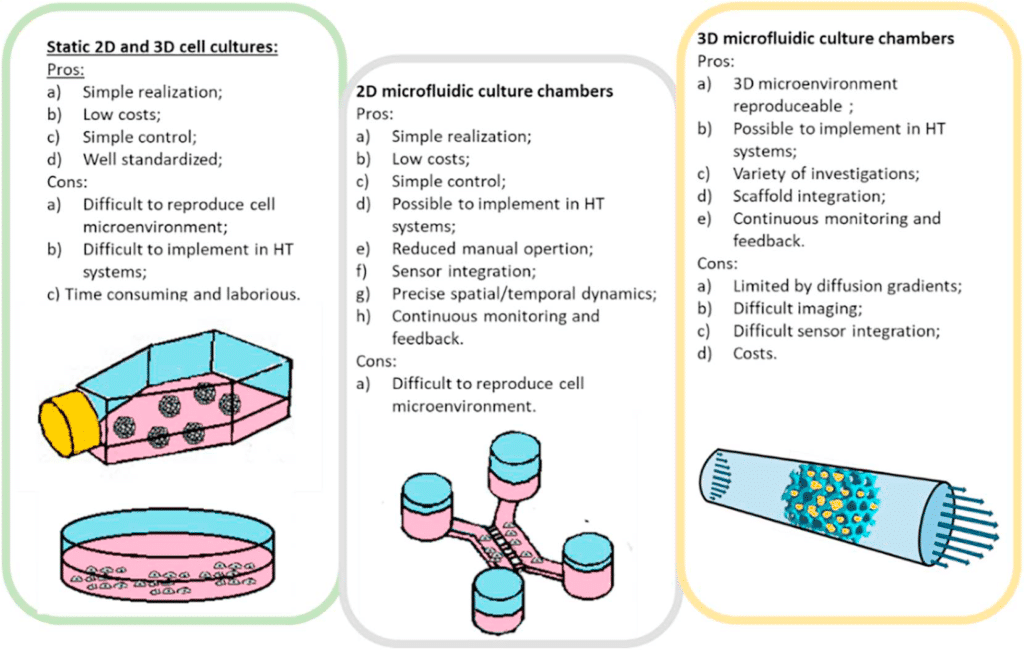
Figure 6: Pro and cons of traditional static cell culture, and 2D and 3D microfluidic cell culture16
Microfluidic chip application model of a tumor microenvironment
The physical microenvironment of tumors is characterized by heterotypic cell interactions and physiological gradients of nutrients, waste products, and oxygen. This tumor microenvironment has a major impact on the biology of cancer cells and their response to chemotherapeutic agents. Despite this, most in vitro cancer research still relies primarily on cells grown in 2D and in isolation in nutrient and oxygen-rich conditions.
Ayuso et al. presented an easy-to-use microfluidic chip application device that can mimic the three-dimensional architecture of multicellular spheroids, while at the same time generating a visible, live “tumor slice” that allows easy monitoring of cells in different regions of the microenvironment in real-time as well as their response to different drugs17 (figure 7).
In this application of microfluidic chip setup, tumor cell behavior in different regions of the microdevice was studied and analyzed in conjugation with measurements of hypoxia and glucose concentrations across the device. The differential cellular response to several well-known drugs in different parts of the microdevice emphasizes the potential of this technology for analyzing the impact of microenvironmental parameters on drug response.
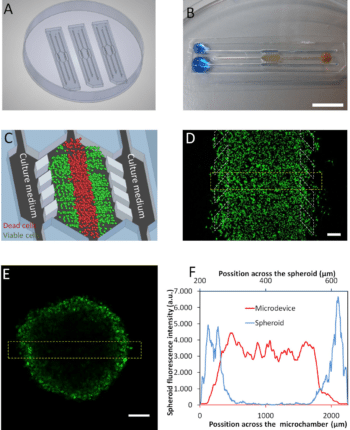
Figure 7: Microdevice operation of the tumor microenvironment.
The figure presents microdevices in a Petri dish containing a central culture chamber and 6 channels. To better understand how the chip works, picture B shows one microdevice filled with (yellowish) collagen hydrogel flowing to the microchamber from the right middle channel and blue-colored water perfused through the two lateral microchannels.
In experimental conditions, the culture medium perfused through the lateral microchannels provides nutrients and oxygen creating physiological gradients across the device. Cells near the ‘surrogate’ blood vessels are viable, whereas oxygen-poor cells in the center of device start to die creating a ‘necrotic core’ similar to the necrotic regions of tumors. It is possible to monitor cells with fluorescent dye in microdevice17 (picture D).
Application notes & expertises:
- Assess Cell Proliferation Using Pressure as a Tool
- Creating a Microfluidic Cancer-on-Chip Platform
- Cancer Cell Analysis Made Easy with Aria: cell Capture and Labeling
- Passive and active mechanical stimulation in microfluidic systems
- Mimicking in-vivo environments: biochemical and biomechanical stimulation
III. Organ on a chip, a cutting edge application of microfluidic chip
Many efforts are devoted to the development of cancer metastasis models that can help in understanding the disease and the development of innovative therapeutic strategies. Current in vitro and in vivo cancer models are incapable of satisfactorily predicting the outcome of various clinical treatments on patients18. Therefore, new application of microfluidic chip methods and approaches for drug discovery and health research are being developed. The concept of mimicking the organ-level function of human physiology or disease using cells inside a microfluidic chip application setup was first published in 2004. In 2010 that the term organ-on-a-chip (OOAC) was invented by Ingber, et. al., who developed a microfluidic chip model to capture organ-level functions of the human lung19.
Microfluidic chip applications enable one the unique ability to control the cellular microenvironment with high spatiotemporal precision and to present cells with mechanical and biochemical signals in a more physiologically relevant context19. The manipulation of the micro-liter volumes of liquids has made these models a platform where scaling, and dynamic crosstalk between cells can be achieved. Microfluidic chips can now use geometries and structures to permit the use of physiological length scales, concentration gradients, and the mechanical forces generated by fluid flow to mimic the in vivo microenvironment experienced by cells20.
These biomimetic applications platforms overcome many drawbacks encountered with conventional tissue culture models. OOAC engineering microfluidic chip application has attracted enormous interest and attention from the pharmaceutical industry, regulatory agencies, and even national defense agencies. This is demonstrated by the increase of OOAC research papers and by the emergence of at least 28 organ-on-a-chip companies in less than seven years21.
A microfluidic chip design to reconstitute organ-level lung functions
To demonstrate that it is possible to engineer a microsystem that replicates the complex physiological functionality of living organs, Huh et al. developed a multifunctional microdevice that reproduces key structural, functional, and mechanical properties of the human alveolar-capillary interface, which is the fundamental functional unit of the living lung19. The microfluidic chip application device consisted of compartmentalized PDMS microchannels to form an alveolar-capillary barrier on a thin, porous, flexible PDMS membrane coated with the extracellular matrix and human alveolar epithelial cells and human pulmonary microvascular endothelial cells is cultured on opposite sides of the membrane (figure 8).
In fact, the device recreates physiological breathing movements (shown in Figure 8.B) by applying vacuum to the side chambers and causing mechanical stretching of the PDMS membrane forming the alveolar-capillary barrier. The device is made of 3 PDMS layers that are bonded to form two sets of three parallel microchannels. (Figure 8.C) PDMS etchant is flowed through the side channels in order to form the two large side chambers. (Figure 8.D). Figure 8.E represents an image of an actual lung-on-a-chip microfluidic device19.
To put it in a nutshell, air is subsequently introduced into the compartment to create an air-liquid interface to mimic the lining of the alveolar air space19 .
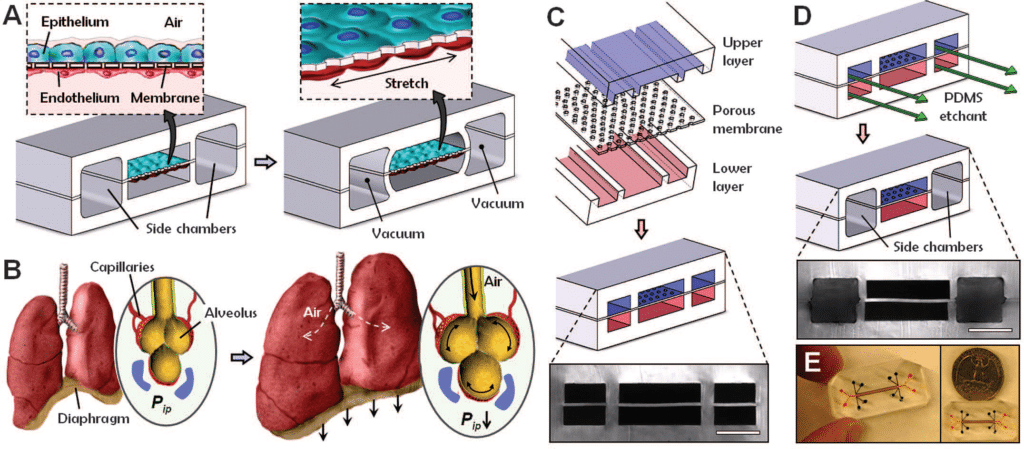
Figure 8: Biologically inspired design of a human breathing lung-on-a-chip microdevice.
Using this microfluidic chip application platform, the authors demonstrated that breathing motions, simulated by the organ-on-chip platform, might greatly accentuate the proinflammatory activities of silica nanoparticles and contribute substantially to the development of acute lung inflammation19. This behavior could not be determined using existing in vitro models.
Application notes & expertises:
- Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications
- Long-term fluid recirculation system for Organ-on-a-Chip applications
- CNRS/UTC: study of a liver-on-a-chip model
- Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions
- Cartilage-on-a-chip, an example of complex mechanical stimulation using Fluigent’s technology
- Creating a Microfluidic Cancer-on-Chip Platform using Fluigent’s High Throughput Cell Perfusion Pack
IV. Particle and cell sorting applications using cell sorter chips
Efficiently isolating and organizing cells from complex mixtures is a crucial task in various fields such as biology, biotechnology, and medicine. Microfluidic chips play a pivotal role in this process, commonly employed to enrich or purify cell samples, thereby enhancing efficiency in research and development22.
Traditionally, optical methods like FACS (Fluorescent Activated Cell Sorting) are used for cell detection. FACS utilizes a laser beam, with the scattered light providing characteristic information about cells and their components. Automated and robust, FACS platform has been a gold standard for cell sorting. However, current commercial platforms face limitations in sample throughput and processing speeds, posing challenges for generating clinical-scale samples.
In contrast, platforms with microfluidic chip offer affordability, simplicity, and a smaller footprint. These chips employ various techniques for cell sorting, each with specific speeds and efficiencies. The chip’s dimensions accommodate diverse cell sorters, ranging from large-volume to precise single-cell sorters.
Additionally, microfluidic cell sorting can integrate various fluidic operations within a single chip, making it versatile for lab-on-a-chip applications, diagnostics, and therapeutics. This approach holds promise in both academic and industrial labs.
Cell sorter in microfluidic devices relies on determining specific cell parameters, such as size, shape, density, or surface markers. Heterogeneous cell solutions are injected into the microfluidic chip, where cells with different properties experience varying forces, leading to their separation into different streamlines and exits.
Microfluidic cell sorting can be categorized into three categories:
- Fluorescent label-based,
- Bead-based
- Label-free cell sorting22
Label-free sorting is perhaps the most studied and comprises both active systems (relying on external fields for sorting) and passive systems that don’t use fluorescent labels or beads. Instead, these methods leverage inherent differences in cellular morphology between cell groups 22.
Inertial microfluidics for continuous particle separation in spiral microchannels
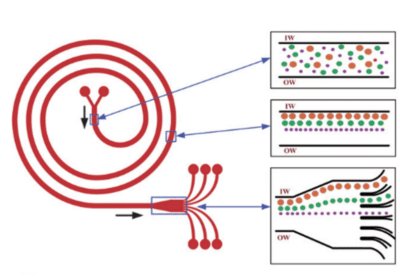
Figure 9: Schematic of the spiral microparticle separator chip.
Kuntaegowdanahalli et al. developed a simple inertial microfluidic device for continuous multi-particle separation, using the Dean-coupled inertial migration principle in spiral microchannels.
In this innovative design, dominant inertial forces, combined with the Dean rotational force arising from the microchannel’s curved geometry, cause particles to settle at a single equilibrium position near the inner microchannel wall. The specific position of particle equilibrium is determined by the ratio of inertial lift to Dean drag forces.
The researchers applied this principle to create a spiral lab-on-a-chip, showcasing size-dependent particle focusing at distinct equilibrium positions across the microchannel cross-section from a multi-particle mixture23. Randomly dispersed particles equilibrate at different positions along the inner wall of the spiral microchannel, influenced by lift and drag forces (see Figure 9 – insert 2). As the particles reach the end of the spiral, they align and separate into different channels (see Figure 9 – insert 3).
A notable advantage of this microfluidic chip application is its high throughput, reaching 1.5 mL/min, achieved without the need for sheath flow or sequential cell manipulation. This feature is particularly beneficial for processing native biological fluids and applications in flow cytometry.
Application notes & expertises:
V. Micromixers an application using microfluidic chips
Micromixers play a crucial role in lab-on-chip devices for various microfluidic chip applications, including drug delivery, sequencing, amplification, and biochemical reactions. They can be broadly classified into two categories based on the actuation method: passive and active.
Passive mixing relies on the microfluidic chip’s geometry and fluid properties without external sources. In laminar flow, typical in microfluidics, mixing primarily occurs through diffusion. This property allows for precise tuning of mixing by employing lamination, where two or more liquids flow in parallel, enabling diffusion to take place. For emproved and faster mixing, chaotic advection can be induced by modifying the microfluidic chip’s geometry, altering channel shapes for splitting, folding, stretching, or disrupting fluid flow.
On the other hand, active mixing involves external perturbation. Various methods are employed in the microfluidic chip application field for active mixing. Dielectrophoresis mixing uses an electric field to move particles toward or away from an electrode, creating chaotic advection. Acoustic wave energy can also mix fluids by generating strong acoustic waves that interfere with each other, creating advection 24. Additionally, adjusting the microfluidic chamber temperature can enhance mixing, as the diffusion coefficient of a liquid is temperature-dependent25.
Submillisecond organic synthesis using a serpentine-shaped microfluidic chip application
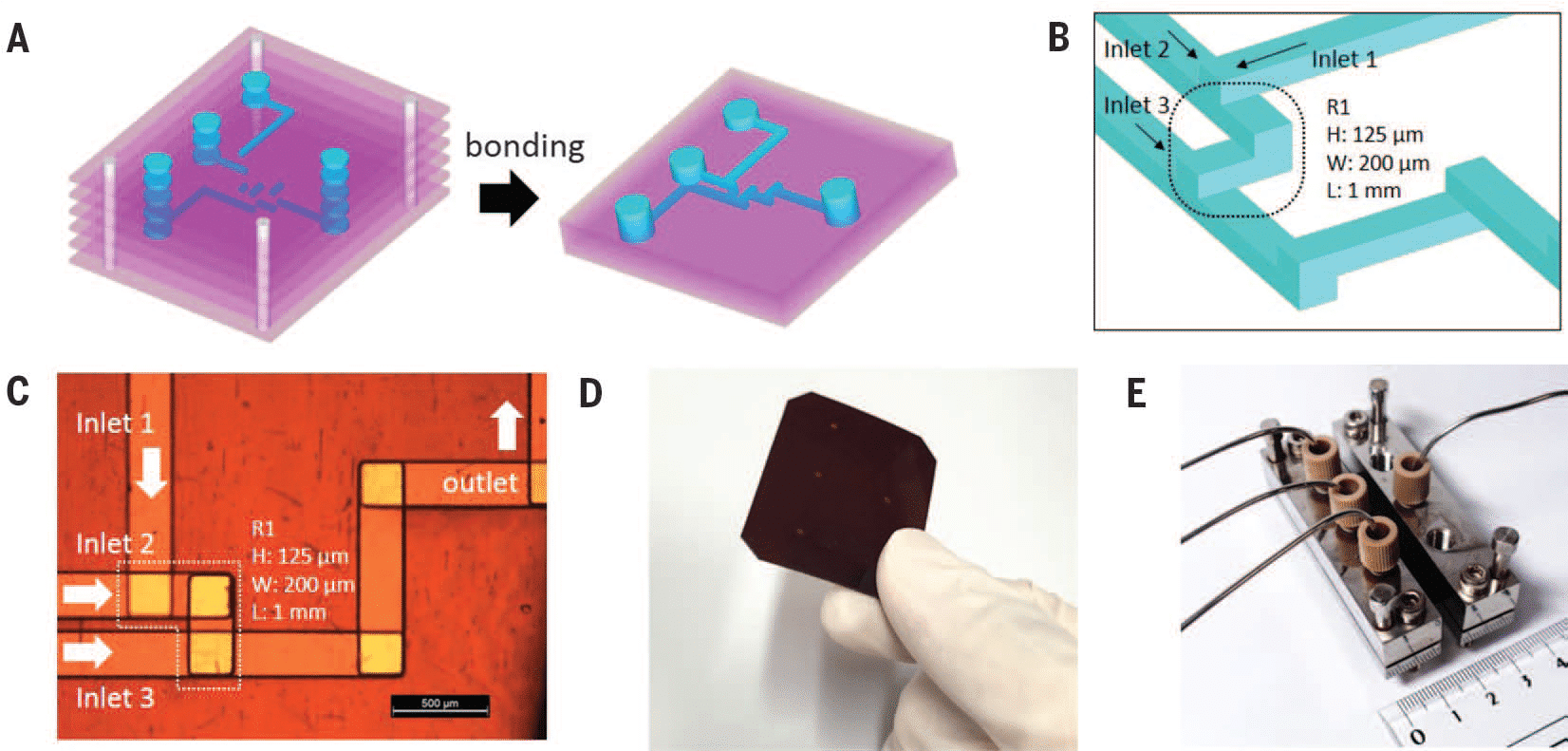
Figure 10: Serpentine-shaped microfluidic chip
In chemical synthesis, it is important to explore the synthetic pathways of an intermediate. To fully observe these pathways, control over its lifetime and mixing time is required.
The reaction mixture had to be well-mixed within the lifetime of the reactive intermediate. An efficient means of prolonging the lifetime is to lower the reaction temperature (-78°C to -100 °C), above the melting point of many organic solvents. Using microfluidic devices, mixing can be extremely fast, with mixing times unattainable by batches26. However, mixing time is increased at low temperatures, as the solvent viscosity exponentially increases.
To circumvent this issue, the authors used a three-dimensional serpentine-shaped microfluidic chip, allowing improved mixing by chaotic advection. The conceptual scheme of the 3D serpentine microchannel fabricated by lamination is represented in figure 10.A.
The figure 10.B represents a detailed scheme for a nanoliter reaction space of rectangular 3D serpentine channels with three inlets and the optical image showing from the top the nanoliter reaction space schematized in B. The optical images of respectively the chip reactor module and assembly are illustrated in Figure 10.D and E.
The utilization of this platform for the application enabled submillisecond mixing.
VI. Microvalves and microfluidic chip applications with reduced internal volume
In the past 10 years, efforts have been devoted to the development of microfluidic platforms capable of performing several assays using programmable fluidic operations within an array of microvalves. Similar to programmable logic circuits where multiple electronic computing routines are executed on a single microdevice, programmable microfluidic platforms have been implemented27, allowing one to perform fluidic operations such as mixing, sampling, washing, and reacting automatically within a single microfluidic chip by modifying the sequence/order of fluidic operations using the software.
The primary advantage of microvalves over their macroscale counterparts is the significantly reduced dead volume, which is important in many microfluidic chip applications that require precise flow control at small flow rates28. They are useful in biological and chemical applications, such as quantitative metabolic biomarker and genetic analysis29,30, protein-based biomarker detection31, or small molecule chemical and environmental analysis32. These microfluidic chip and valve application platforms usually consist of a 2D array of microvalves that permit flow regulation, on/off switching and sealing of liquids, gases, or vacuums33.
Several microvalves have been developed using pneumatic, electrokinetic and electrochemical actuators. Among these mechanisms, pneumatic actuation is often recognized as the most reliable method due to the simplicity of fabrication, ease of use, scalability, reliability, and a high degree of accuracy. Pneumatically actuated microvalves utilize the deflection of an elastomer (typically PDMS) membrane to control fluid flow34.
A fully integrated multilayer microfluidic chemical analyze for automated sample processing, labelling, and analysis
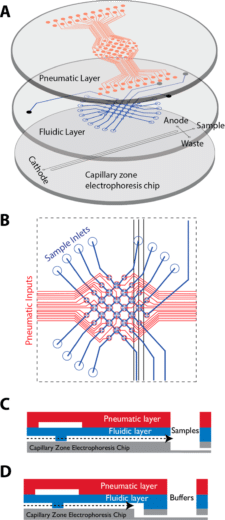
Figure 11: (A) Schematic of the microfluidic automaton integrated with the Capillary Zone Electrophoresis (CZE) analysis chip. (B) Detailed view of cellular microvalve array. (C) and (D) Cross-sectional view of the CZE sample and buffers32.
Capillary zone electrophoresis (CZE), is a powerful tool for chemical analysis and is widely used for environmental monitoring, astrobiology, and biosensing32. CZE assays usually require complex and manual sample processing.
Commercial platforms for automated CZE have been implemented to address this concern, but are expensive, and large, thus hardly field deployable in challenging environments. Microfluidic application chips and devices allow miniaturization, automation, and reduction in sample volume requirements for chemical and biochemical sensing.
Using membrane microvalve technology, it is possible to automate metering, transporting, routing, and mixing operations. Kim et al. introduced a microfluidic platform consisting of pneumatically actuated “lifting gate” microvalves integrated with a glass CZE microchip, providing extremely low dead volumes between components.
All the procedures, including buffer filling, labeling, and dilution, can be automated. The microchip was used to analyze diverse compound classes, such as amino acids, and oxidized biomarker compounds, like aldehydes/ketones and carboxylic acids in less than 30 min.
VII. Wearable microfluidics: an emerging application
An emerging application of microfluidic chip, is the use of microfluidic concepts for wearable device applications36. Here, microfluidics presents several key value propositions. The microstructures store or handle fluids and are the core of the sensing device. Using microchannels of the microfluidic chip, precise liquid amounts can be manipulated, allowing for highly accurate and reliable devices and being useful for bodily fluids that are often secreted or extracted in limited quantities.
Also, wearable microfluidic devices could also stock a specific drug for precise dispensing at controlled intervals. Innovations in flexible microfluidics and electronics have led to numerous applications. Typically, a wearable microfluidic device will collect a fluid, transfer it to a localized site where detection or measurement is performed. Blood and sweat are common analytes as they provide insights into physiological states such as temperature, pH, and hydration36. Wearable microfluidics finds applications in the pharmaceutical, food, sportswear, and cosmetic fields.
A wearable microfluidic device for the capture, storage and colorimetric sensing of sweat
A wearable microfluidic device for the capture, storage and colorimetric sensing of sweat

Figure 11: (A) Schematic of the microfluidic automaton integrated with the Capillary Zone Electrophoresis (CZE) analysis chip.
As mentioned previously, sweat is an analyte of interest because of its rich content of important biomarkers. It is easy to collect compared to blood. In situ quantitative analysis of sweat is of great interest for monitoring physiologic health status (for example, hydration state) and for the diagnosis of disease37.
Existing systems for sweat collection and analysis are confined to laboratories, where standard analytical technologies can be performed. Though highly precise, the analysis is time-consuming and costly.
To address this issue, Kho et al. developed a soft wearable microfluidic system than can directly harvest sweat from pores on the surface of the skin37.
The device routes the sample to different channels and reservoirs for multiparametric sensing of markers of interest, with options for wireless interfaces to external devices for image capture and analysis.
The device can measure total sweat loss, pH, lactate, chloride, and glucose concentrations by colorimetric detection using wireless data transmission. As it is a simple, low-cost, and fast analysis point of care device, it could be used to accumulate data from individual users over time, and this could serve as an analytical approach for interpreting trends in marker concentrations, potentially providing warning signs when performing physical activity.
Conclusion
Since the introduction of microfluidics, the scope of microfluidic chip applications has kept extending over the years. The first applications were focused on analytical chemistry, but today the field of life science and specifically point of care is in the core of microfluidics. We have presented applications where microfluidic chips show great advantages compared to conventional systems. The importance of these applications was illustrated by showing research papers related to these applications. Some important applications of microfluidic chip were introduced here. Microfluidics covers a wide range of applications such as microreactors, bioprinting, fuel cells, and many more.
Expertises
-
Microfluidics Case Studies Creating a Microfluidic Cancer-on-Chip Platform using Fluigent’s High Throughput Cell Perfusion Pack Read more
-
Microfluidic Application Notes Long-term fluid recirculation system for Organ-on-a-Chip applications Read more
-
Microfluidics Case Studies CNRS/UTC: study of a liver-on-a-chip model Read more
-
Microfluidic Application Notes Alginate Microcapsule Synthesis Read more
-
Microfluidic Application Notes Agarose Microcapsules Synthesis Read more
-
Microfluidic Application Notes PLGA nanoparticle synthesis using 3D microfluidic hydrodynamic focusing Read more
-
Microfluidic Application Notes Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions Read more
-
Microfluidic Application Notes PLGA microcapsules synthesis Read more
-
Microfluidic Application Notes Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications Read more
-
Microfluidic Application Notes Double Emulsion Generation Read more
-
Microfluidic Application Notes Microfluidic Chitosan Microcapsules Production Read more
-
Microfluidic Application Notes Alginate Microbeads Production Read more
-
Microfluidic Application Notes Single cell sorter microfluidic platform Read more
-
Microfluidic Application Notes Assess Cell Proliferation Using Pressure as a Tool Read more
-
Microfluidic Application Notes Cartilage-on-a-chip, an example of complex mechanical stimulation using Fluigent’s technology Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
References
- Seemann, R., Brinkmann, M., Pfohl, T. & Herminghaus, S. Droplet based microfluidics. Reports Prog. Phys. 75, (2012).
- Paquin, F., Rivnay, J., Salleo, A., Stingelin, N. & Silva, C. Droplet Control Technologies for Microfluidic High Throughput Screening (µHTS). Muhsincan Sesen,a Tuncay Alan,a and Adrian Neild∗a 10715–10722 (2017) doi:10.1039/b000000x.
- Galas, J. C., Bartolo, D. & Studer, V. Active connectors for microfluidic drops on demand. New J. Phys. 11, (2009).
- Jullien, M.-C., Tsang Mui Ching, M.-J., Cohen, C., Menetrier, L. & Tabeling, P. Droplet break in a low capillary T-junction. in 19th Mechanical French Congress (AFM, Maison de la Mécanique, 39/41 rue Louis Blanc-92400 Courbevoie, 2009).
- Yu, L., Chen, M. C. W. & Cheung, K. C. 2010 Droplet-based microfluidic system for multicellular tumor spheroid formation and anticancer drug testing. Lab Chip 10, 2424–2432 (2010).
- N.Shembekar, C.Chaipan, R. U. & C. A. M. 2016 Droplet-based microfluidics in drug discovery, transcriptomics and high-throughput molecular genetics. Lab Chip (2016) doi:10.1039/C6LC00249H.
- Shang, L., Cheng, Y. & Zhao, Y. Emerging Droplet Microfluidics. Chem. Rev. 117, 7964–8040 (2017).
- Suea-Ngam, A., Howes, P. D., Srisa-Art, M. & Demello, A. J. Droplet microfluidics: From proof-of-concept to real-world utility? Chem. Commun. 55, 9895–9903 (2019).
- Vladisavljević, G. T., Al Nuumani, R. & Nabavi, S. A. Microfluidic production of multiple emulsions. Micromachines 8, (2017).
- Soppimath, K. S. & Aminabhavi, T. M. Ethyl acetate as a dispersing solvent in the production of poly(DL-lactide-co-glycolide) microspheres: Effect of process parameters and polymer type. J. Microencapsul. 19, 281–292 (2002).
- Qi, F., Wu, J., Li, H. & Ma, G. Recent research and development of PLGA / PLA microspheres / nanoparticles : A review in scienti fi c and industrial aspects. (2018).
- Anderson, J. M. & Shive, M. S. Biodegradation and biocompatibility of PLA and PLGA microspheres. Adv. Drug Deliv. Rev. 64, 72–82 (2012).
- Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
- Halldorsson, S., Lucumi, E., Gómez-Sjöberg, R. & Fleming, R. M. T. Advantages and challenges of microfluidic cell culture in polydimethylsiloxane devices. Biosens. Bioelectron. 63, 218–231 (2015).
- Yeo, L. Y., Chang, H. C., Chan, P. P. Y. & Friend, J. R. Microfluidic devices for bioapplications. Small 7, 12–48 (2011).
- Coluccio, M. L. et al. Microfluidic platforms for cell cultures and investigations. Microelectron. Eng. 208, 14–28 (2019).
- Ayuso, J. M. et al. Development and characterization of a microfluidic model of the tumour microenvironment. Sci. Rep. 6, 1–16 (2016).
- Caballero, D. et al. Organ-on-chip models of cancer metastasis for future personalized medicine: From chip to the patient. Biomaterials 149, 98–115 (2017).
- Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science (80-. ). 328, 1662–1668 (2010).
- Bhise, N. S. et al. Organ-on-a-chip platforms for studying drug delivery systems. J. Control. Release 190, 82–93 (2014).
- Zhang, B., Korolj, A., Lai, B. F. L. & Radisic, M. Advances in organ-on-a-chip engineering. Nat. Rev. Mater. 3, 257–278 (2018).
- C. Wyatt Shields IV, Dr. Catherine D. Reyes, and P. G. P. L. Microfluidic Cell Sorting: A Review of the Advances in the Separation of Cells from Debulking to Rare Cell Isolation. Lab Chip (2015) doi:10.1039/c4lc01246a.
- Kuntaegowdanahalli, S. S., Bhagat, A. A. S., Kumar, G. & Papautsky, I. Inertial microfluidics for continuous particle separation in spiral microchannels. Lab Chip 9, 2973–2980 (2009).
- Vivek, V., Zeng, Y. & Kim, E. S. NOVEL ACOUSTIC-WAVE MICROMIXER.
- Ward, K. & Fan, Z. H. Mixing in microfluidic devices and enhancement methods. J. Micromechanics Microengineering 25, (2015).
- Kim, H. et al. Submillisecond organic synthesis: Outpacing Fries rearrangement through microfluidic rapid mixing. Science (80-. ). 352, 691–694 (2016).
- Thorsen, T., Maerkl, S. J. & Quake, S. R. Microfluidic large-scale integration. Science (80-. ). 298, 580–584 (2002).
- Aditya Aryasomayajula, Pouriya Bayat, Pouya Rezai, P. R. S. Microfluidic Devices and Their Applications. Springer vol. 50 (2017).
- Jensen, E. C., Bhat, B. P. & Mathies, R. A. A digital microfluidic platform for the automation of quantitative biomolecular assays. Lab Chip 10, 685–691 (2010).
- Vincent, M., Xu, Y. & Kong, H. Helicase-dependent isothermal DNA amplification. EMBO Rep. 5, 795–800 (2004).
- Erik C. Jensen, Yong Zeng, Jungkyu Kim, and R. A. M. Microvalve Enabled Digital Microfluidic Systems for High Performance Biochemical and Genetic Analysis. 15, 455–463 (2011).
- Kim, J., Jensen, E. C., Stockton, A. M. & Mathies, R. A. Universal microfluidic automaton for autonomous sample processing: Application to the mars organic analyzer. Anal. Chem. 85, 7682–7688 (2013).
- Oh, K. W. & Ahn, C. H. A review of microvalves. J. Micromechanics Microengineering 16, (2006).
- Kim, J., Stockton, A. M., Jensen, E. C. & Mathies, R. A. Pneumatically actuated microvalve circuits for programmable automation of chemical and biochemical analysis. Lab Chip 16, 812–819 (2016).
- Chin, V. I. et al. Microfabricated platform for studying stem cell fates. Biotechnol. Bioeng. 88, 399–415 (2004).
- Yeo, J. C., Kenry & Lim, C. T. Emergence of microfluidic wearable technologies. Lab Chip 16, 4082–4090 (2016).
- Ahyeon Koh, Daeshik Kang, Yeguang Xue, Seungmin Lee, Rafal M. Pielak, Jeonghyun Kim, Taehwan Hwang, Seunghwan Min,1 Anthony Banks, Philippe Bastien, Megan C. Manco, Liang Wang, Kaitlyn R. Ammann, Kyung-In Jang, Phillip Won Seungyong Han, Roozbeh Ghaffari, J. A. R. A soft, wearable microfluidic device for the capture, storage, and colorimetric sensing of sweat. Sci. Transl. Med. 26, 39–46 (2017).
The Lamprou Lab (Queen’s University Belfast)
The Lamprou lab, affiliated with Queen’s University Belfast, specializes in three main areas: nanoparticles for imaging and therapy, lab-on-a-chip technology, and therapeutic implants. Their interdisciplinary approach has driven innovation in healthcare since 2012, developing emerging technologies and novel drug delivery devices. The lab is led by Professor Dimitrios Lamprou, a leading expert in pharmaceutical technologies known for his significant contributions to 3D printing, microfluidics, and nanofibers, with over 150 peer-reviewed publications.


What are engineered nanoparticles?
Nanotechnology gained prominence after Richard P. Feynman’s 1959 famous lecture, “There’s plenty of room at the bottom”.1 However, the use of nanoparticles can be traced back to ancient times, as seen in the Romans’ incorporation of nanoparticles into glass manufacturing in the fourth century AD. The Lycurgus cup, an artifact from this period, notably displayed distinctive color changes under various lighting conditions due to the integration of nano-glass particles.2


In modern times, nanotechnology has developed into a comprehensive scientific discipline with diverse applications spanning multiple industries. From water purification and information technologies to drug development, environmental solutions, and the creation of robust yet lightweight materials, nanotechnology has emerged as a pivotal player.3,4
The fundamental units of nanotechnology are nanoparticles, defined as small particles ranging from 10 nm to 1000 nm in size. Engineered nanoparticles, specifically designed with dimensions under 100 nm, play a crucial role in manipulating materials at the molecular and atomic levels, demonstrating significant chemical, structural, electrical, biological and mechanical characteristics. They are classified into categories including ceramic, carbon-based, semiconductor, metal, lipid-based and polymeric nanoparticles.
What makes engineered nanoparticles promising in nanomedicine?
Engineered nanoparticles hold promise for various nanotechnology applications in medicine, including in-vivo and in-vitro diagnosis, drug delivery, and production of biocompatible materials. They are characterized by their high mass-to-surface area ratio, ability to adsorb and carry compounds, and quantum properties.5,6
These characteristics can benefit the drug delivery field, as its primary objective is to enhance the specificity of drug targeting, improve safety and biocompatibility, and reduce toxicity while maintaining therapeutic effects. Nanoparticles with dimensions less than 100 nm are considered excellent drug carriers due to their unique biological and physiological properties, allowing them to cross tissue and cell barriers effectively.7
Engineered nanoparticles also exhibit a higher likelihood of cellular uptake due to their larger surface area, enabling increased protein loading. The interaction of NPs with biological substances results in the formation of a “protein corona,” enhancing nanocarriers’ uptake by the reticuloendothelial system. This protein corona can serve as a functional carrier for targeted drug delivery, improving poorly soluble drug uptake, directing drugs to specific locations, and enhancing drug bioavailability.8
Overall, engineered nanoparticle applications in nanomedicine have been progressing recently, particularly in controlled drug delivery, nucleic acid-based treatment, cancer cell targeting, angiogenesis inhibition, and inflammation control.
How are engineered nanoparticles traditionally produced
The success of nanoparticles as drug carriers and in nanomedicine applications depends on a number of crucial factors, including NP fabrication strategies, physical properties, drug loading efficiencies, drug release potential, and especially the carrier’s toxicity.
Lipid-based nanoparticles exhibit low toxicity in in-vivo experiments. They can carry both hydrophilic and hydrophobic molecules, leading to prolonged half-life and controlled drug release. Here, we focus on two main classes of lipid-based nanoparticles: liposomes and solid lipid nanoparticles.
Liposomes
A liposome is a microsphere lipid constructed from one or multiple phospholipid bilayers, closely mirroring the structure of cell membranes. The liposome preparation process consists of three main stages: preparing aqueous and lipid phases, primary processing with lipids, and optional secondary processing steps. Most methods involve dissolving phospholipids in an organic solvent, with subsequent removal of the solvent through evaporation—a critical step in liposome formation. Two straightforward methods of liposome synthesis are film hydration and solvent injection.
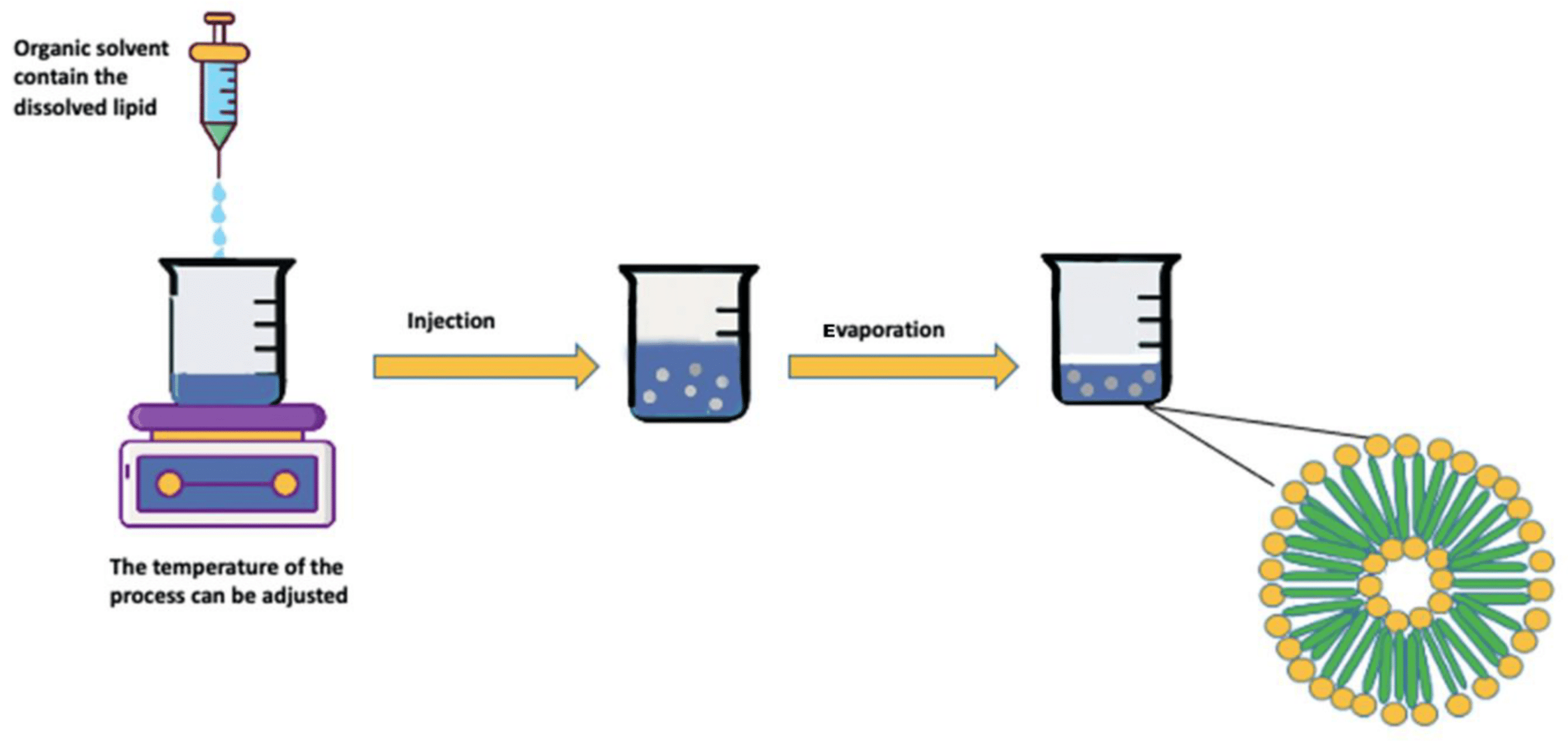
Liposomes generated through film hydration tend to be polydisperse. Parameters like the duration of rotary evaporation, mixing speed, and temperature after hydration affect liposome quality, emphasizing the need for careful monitoring.10,11
The major factors to consider for use of the solvent injection method are the temperature during injection and the injection rate. These factors will affect the size, shape, and polydispersity of the liposomes produced. The solvent injection method involves certain challenges, with continuous exposure of therapeutic substances to high temperatures and organic solvents affecting liposomal product stability and safety.
This results in high polydispersity and a non-homogeneous formulation.12
Solid Lipid Nanoparticles
Solid lipid nanoparticles (SLN) have emerged as a promising alternative to other lipid formulations like liposomes or polymeric nanoparticles. These spherical colloidal particles, ranging from 10 to 1000 nm in size, offer controlled drug release due to limited drug mobility in the solid lipid. SLNs provide advantages such as targeted drug delivery, controlled release, increased stability, scalability in production, and avoidance of organic solvents. Comprising a solid lipid core in an aqueous medium with a surfactant, SLNs use different lipids (steroids, fatty acids, triglycerides, etc.) and require stabilizing agents like surfactants or emulsifiers. The drug insertion process depends on drug hydrophobicity, solid lipid category, and polymeric alterations in the lipid. Production techniques include high-pressure homogenization, solvent evaporation, ultrasonication, hot homogenization, microemulsion, and others.13,14
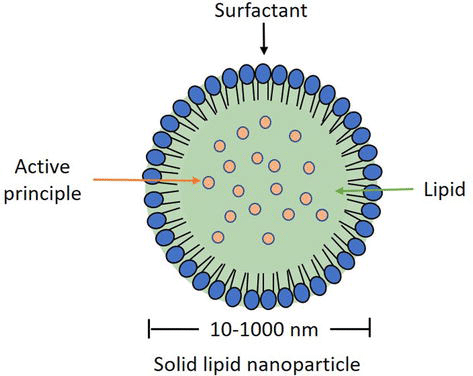
Table: comparison of the different manufacturing methods used for polymeric NPs and lipid-based NPs
| Nanoparticles Type | Manufacturing Method | Advantages | Disadvantages |
|---|---|---|---|
| Lipid formulation | Film hydration | – Established method – Understood method | – High consuming of the organic solvents – High PDI – Lack of reproducibility – Need for additional downsizing step – Difficulties in scaling-up |
| Lipid formulation | Solvent injection | – Simple and fast – Scaling-up possibility | – Exposing to organic solvent – High PDI – Stability problems |
| Lipid formulation | Extrusion | – Uniform and homogenous formulation | – Possible clogging of the membrane pores. – Difficulties in scaling-up |
| Lipid formulation | High pressure homogenization | – Scaling-up possibility – Uniform formulation | – High energy consumption – Multiple steps – Bulky system |
| Lipid formulation | Microemulsion | – Small particle size – Homogenous formulation | – Difficulty in removing the excess water – Use high concentration of surfactants |
| Nanoparticles Type | Manufacturing Method | Advantages | Disadvantages |
|---|---|---|---|
| Polymeric | Emulsification-salting out | – Avoids surfactants and chlorinated solvents | – Need for purification steps – Encapsulate lipophilic drugs only |
| Polymeric | Emulsification solvent diffusion | – Scaling-up possibility – Batch-to-batch reproducibility | – The possible diffusion of the hydrophilic drug into the aqueous phase – The need to eliminate high volume of aqueous phase from the colloidal dispersion |
| Polymeric | Emulsification- evaporation | – Simple and versatile | – Risk of nanodroplets coalescence during the evaporation process – Time consuming |
| Polymeric | Dialysis | – Effective and simple method – Produce polymeric nanoparticles with narrow distribution | – Time consuming – Use of high amount of dialyzing medium, which stimulate the premature release of NPs content |
| Polymeric | Nonparticipation | – Simple and established method – Use low concentrations of surfactant | – Restricted for lipophilic drugs – Low polymer concentration obtained |
How does microfluidics enhance the properties of engineered nanoparticles
The principal innovation of microfluidics is the ability to transfer the traditional bulk technique to microscale fluidic chips. Solvents can be mixed within microchannels by a pumping system with continuous laminar flow. This type of flow offers high mixing quality and enhances the performance of microscale devices. The ability to adjust the flow rate ratio (FRR) and total flow rate (TFR) allows for continuous production of monodisperse and homogenous engineered nanoparticles.
Fluigent’s pressure-driven controllers and flow sensors enable this high stability and fast response in fluid flow, which is highly challenging to achieve with traditional pumping methods like peristaltic or syringe pumps. Overall, this method offers high reproducibility and low batch-to-batch variations. In addition, the method’s versatility makes the encapsulation process faster while keeping encapsulation efficiency high.
Microfluidics offers a fast, simple, single-step technique for liposome manufacturing, but challenges for large-scale production currently hinder its wide-scale implementation due to high costs.15,16
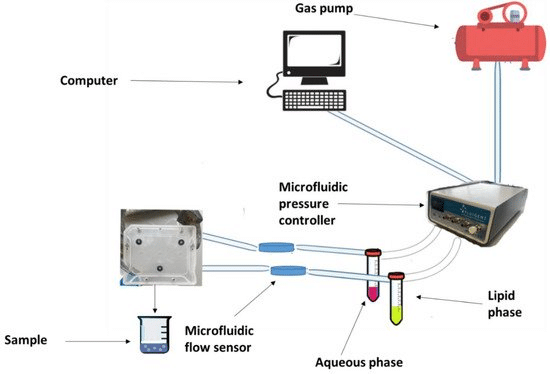
Most microfluidics-based solid lipid nanoparticle manufacturing follows a similar procedure involving dissolving lipids and drugs in an organic solvent, which is then introduced into the microfluidic device alongside an aqueous phase with a surfactant. Control over lipid-to-drug concentration, flow rate, and velocity influences the final nanocarrier characteristics.
Solid lipid nanoparticles produced by microfluidic processes have smaller particle sizes, better homogeneity, and higher encapsulation efficiency compared to bulk methods. In addition, cytotoxic studies demonstrate potent anti-proliferative effects of microfluidic SLNs in cancer cell lines.
However, research on microfluidic-produced SLNs is limited, and challenges exist, particularly with regard to the material used for microfluidic chips, with PDMS-based chips being sensitive to organic solutions.17
Conclusion
In this paper highlight, we presented the promise and challenges of microfluidics technology for the design and formulation of nanomedicines. While traditional methods face limitations in producing small engineered nanoparticles with desirable characteristics, microfluidic systems offer a one-step, controllable process with improved outcomes.
Comparative studies have shown that nanocarriers produced by microfluidics exhibit superior properties. Future advancements in microfluidic systems, combined with complementary tools like process analytical technology and molecular imaging technologies, are expected to optimize NP production and expand their medical applications.
Microfluidics holds promise for shaping the future of engineered nanoparticle research and development.
Related Products
Expertises & Resources
-
Expert Reviews: Basics of Microfluidics Giant Unilamellar Vesicles (GUVs) Production using Microfluidics Read more
-
Microfluidics Article Reviews Solid lipid nanoparticles for biologics and drug encapsulation Read more
-
Microfluidics Article Reviews A mRNA encapsulation platform integrating Fluigent’s FlowEZ Read more
-
Expert Reviews: Basics of Microfluidics Microfluidics for vaccine development Read more
-
Microfluidic Application Notes Liposome Nanoparticle Synthesis Read more
-
Expert Reviews: Basics of Microfluidics Flow control for droplet generation using syringe pumps and pressure-based flow controllers Read more
References
(1) Shirai, Y.; Osgood, A.J.; Zhao, Y.; Yao, Y.; Saudan, L.; Yang, H.; Yu-Hung, C.; Alemany, L.B.; Sasaki, T.; Morin, J.-F. Surface-rolling molecules. J. Am. Chem. Soc. 2006, 128, 4854–4864.
(2) Bayda, S.; Adeel, M.; Tuccinardi, T.; Cordani, M.; Rizzolio, F. The History of Nanoscience and Nanotechnology: From Chemical-Physical Applications to Nanomedicine. Molecules 2019, 25, 112. Schoenmaker L, et al. mRNA-lipid nanoparticle COVID-19 vaccines: structure and stability. Int Pharm. 2021;601: 120586 .
(3) Grobert, N.; Hutton, D. Nanoscience and nanotechnologies: Opportunities and uncertainties. Lond. R. Soc. R. Acad. Eng. Rep.2004, 46, 618.
(4) Thiruvengadam, M.; Rajakumar, G.; Chung, I.M. Nanotechnology: Current uses and future applications in the food industry.3 Biotech 2018, 8, 74.
(5) Duncan, R. The dawning era of polymer therapeutics. Nat. Rev. Drug Discov. 2003, 2, 347–360.
(6) De Jong, W.H.; Borm, P.J. Drug delivery and nanoparticles:applications and hazards. Int. J. Nanomed. 2008, 3, 133–149.
(7) LaVan, D.A.; McGuire, T.; Langer, R. Small-scale systems for in vivo drug delivery. Nat. Biotechnol. 2003, 21, 1184–1191.
(8) Nel, A.E.; Mädler, L.; Velegol, D.; Xia, T.; Hoek, E.M.; Somasundaran, P.; Klaessig, F.; Castranova, V.; Thompson, M. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 2009, 8, 543–557.
(9) Mitchell, M.J., Billingsley, M.M., Haley, R.M. et al. Engineering precision nanoparticles for drug delivery. Nat Rev Drug Discov 20, 101–124 (2021).
(10) Jaradat, E.; Weaver, E.; Meziane, A.; Lamprou, D. A. Microfluidics Technology for the Design and Formulation of Nanomedicines. Nanomaterials 2021, 11 (12), 3440.
(11) Alam, S.; Mattern-Schain, S.; Best, M. Targeting and triggered release using lipid-based supramolecular assemblies as medicinal nanocarriers. In Comprehensive Supramolecular Chemistry II; Elsevier: Oxford, UK, 2017; pp. 329–364.
(12) Maherani, B.; Arab-Tehrany, E.; Mozafari, M.R.; Gaiani, C.; Linder, M. Liposomes: A review of manufacturing techniques and targeting strategies. Curr. Nanosci. 2011, 7, 436–452.
(13) Mehnert, W.; Mäder, K. Solid lipid nanoparticles: Production, characterization and applications. Adv. Drug Deliv. Rev. 2001, 47,165–196.
(14) Uner, M.; Yener, G. Importance of solid lipid nanoparticles (SLN) in various administration routes and future perspectives. Int. J.Nanomed. 2007, 2, 289–300.
(15) Weaver, E.; Uddin, S.; Cole, D.K.; Hooker, A.; Lamprou, D.A. The Present and Future Role of Microfluidics for Protein and Peptide-Based Therapeutics and Diagnostics. Appl. Sci. 2021, 11, 4109.
(16) Guimarães Sá Correia, M.; Briuglia, M.L.; Niosi, F.; Lamprou, D.A. Microfluidic manufacturing of phospholipid nanoparticles: Stability, encapsulation efficacy, and drug release. Int. J. Pharm. 2017, 516, 91–99.
(17) Arduino, I.; Liu, Z.; Rahikkala, A.; Figueiredo, P.; Correia, A.; Cutrignelli, A.; Denora, N.; Santos, H.A. Preparation of cetylpalmitate-based PEGylated solid lipid nanoparticles by microfluidic technique. Acta Biomater. 2021, 121, 566–578.
Organ-on-a-chip (OOAC) is the concept of mimicking the organ-level function of human physiology or disease using cells inside a microfluidic chip. Microfluidics provides the unique ability to control the cellular microenvironment with high spatiotemporal precision and to present cells with mechanical and biochemical signals in a more physiologically relevant context. The ability to manipulate micro-liter volumes of liquids has made these models a platform where scaling and dynamic crosstalk between cells can be achieved.
Microfluidic chips can now use geometries and structures to permit the use of physiological length scales, concentration gradients, and the mechanical forces generated by fluid flow to mimic the in vivo microenvironment experienced by cells. These biomimetic platforms overcome many of the drawbacks encountered with conventional tissue culture models.
Applications of OOAC cell culture
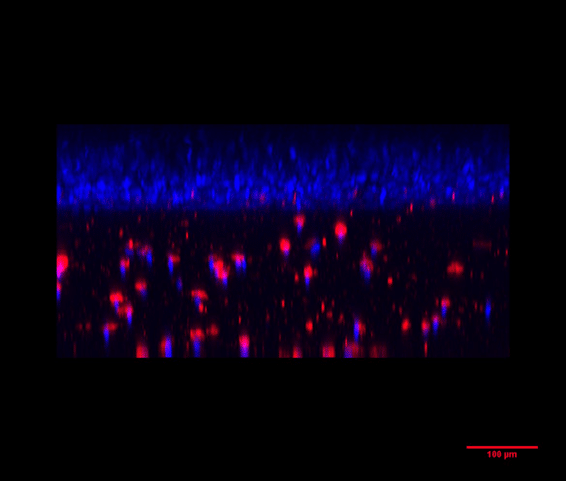
Therapy development
Organ-on-chip cell culture platforms have proven potential in providing tremendous flexibility and robustness in drug screening and development by employing engineering techniques and materials. More importantly, there is a clear upward trend in studies that utilize human-induced pluripotent stem cells (hiPSC) to develop personalized tissue or organ models. For this purpose, the use of cell culture chips allows users to study complex culture configurations by joining a culture well with a microfluidic channel via a porous membrane. This is the optimal device for Air Liquid Interface (ALI) culture, endothelium/epithelium barrier and crosstalk studies.

Drug discovery
The development of emerging in-vitro tissue culture platforms can be useful for predicting the human response to new compounds. Recently, several in-vitro tissue-like microsystems, also known as “organ-on-a-chip studies”, have emerged to provide new tools for better evaluating the effects of various chemicals on human tissue.
Organ-on-chip cell culture models can therefore be used for accurate prediction and mechanistic investigation of dose-limiting human toxicities of prospective drugs, as well as for the exploration of new therapeutic approaches to mitigate the observed toxic effects. In the drug discovery pipeline, predictions made by these models could inform and facilitate early efforts to identify, modify and optimize lead compounds, thereby developing safer drugs with an increased likelihood of success in clinical trials.
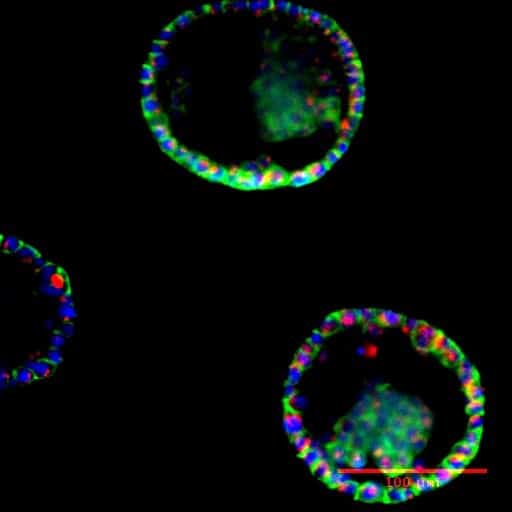
Regenerative medicine
The development of emerging in-vitro tissue culture platforms can be useful for predicting the human response to new compounds. Recently, several in-vitro tissue-like microsystems, also known as ‘organ-on-a-chip studies’, have emerged to provide new tools for better evaluating the effects of various chemicals on human tissue.
Organ-on-chip cell culture models can therefore be used for accurate prediction and mechanistic investigation of dose-limiting human toxicities of prospective drugs, as well as for the exploration of new therapeutic approaches to mitigate the observed toxic effects. In the drug discovery pipeline, predictions made by these models could inform and facilitate early efforts to identify, modify and optimize lead compounds, thereby developing safer drugs with an increased likelihood of success in clinical trials.
Omi, the new automated organ-on-chip platform
Discover Omi, the new automated platform for OOAC applications developed by Fluigent. Equipped with state-of-the-art technologies, this platform will enable you to carry out any perfusion protocol. It has the ability to customize and automate any protocol, including simple perfusion, recirculation, sampling and injection. It meets the needs of beginners in organ-on-chip cell culture research and advanced organ-on-chip researchers looking for automation and reproducibility.
This versatile, automated organ-on-a-chip platform can perform long-term OOAC cell cultures under flow to control shear stress conditions. Its two-hour battery life and WiFi connectivity ensure easy, intuitive control. It can be easily transported from incubator to microscope for live cell imaging while maintaining cell perfusion under battery power. You can also monitor your experiment from anywhere using the Omi app.
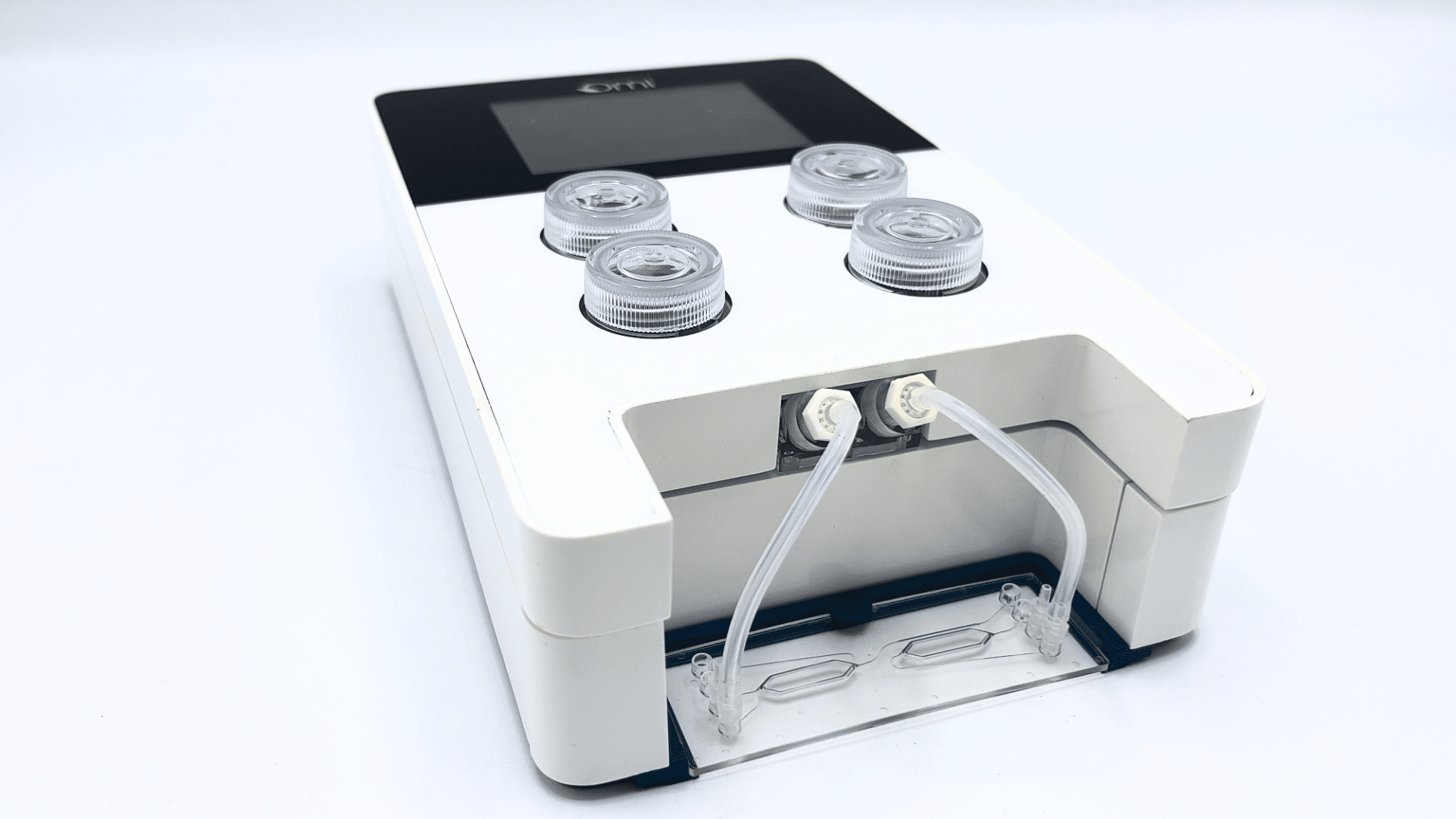

Towards the next generation of organ-on-a-chip cell culture platforms
Fluigent and Beonchip are partnering to offer a complete solution for organ-on-chip cell culture.
This webinar will first introduce Beonchip, their chips and the numerous applications that can be performed with them.
The second part will focus on flow control systems. Although often considered secondary, flow control is as important as chip design, as cells are highly sensitive to mechanical forces. Results demonstrate how cells are affected by peristaltic pumps compared to pressure control systems.
Organ-on-chip research is an emerging field that offers substantial benefits compared to conventional cell culture. In many labs, considerable effort is put into choosing the right chip design, but the impact of flow control is still undetermined.
It is our intent to create awareness of the importance of flow and its effects on studies.
Read our expertise page to learn more about the benefits of flow control in cell culture and about our products and those of our partners (Beonchip), and to reach out to our team of experts to find the solution that best meets your needs.
Resources
-
Microfluidic Application Notes Long-term fluid recirculation system for Organ-on-a-Chip applications Read more
-
Microfluidics Case Studies CNRS/UTC: study of a liver-on-a-chip model Read more
-
Expert Reviews: Basics of Microfluidics Why Control Shear Stress in Cell Biology? Read more
-
Microfluidic Application Notes Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions Read more
-
Microfluidic Application Notes Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications Read more
-
Microfluidic Application Notes Assess Cell Proliferation Using Pressure as a Tool Read more
-
Microfluidic Application Notes Cartilage-on-a-chip, an example of complex mechanical stimulation using Fluigent’s technology Read more
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Prostate Organoid Culture in Microbeads Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
-
Expert Reviews: Basics of Microfluidics MYOCHIP | H2020 European project Read more
-
Expert Reviews: Basics of Microfluidics Application of microfluidic chip technology Read more
Importance of fluid handling for organ-on-a-chip cell culture
In many labs, considerable effort is put into choosing the right chip design, but the impact of flow control is still undetermined. It is our intent to create awareness of the importance of flow and its effects on one’s studies.
Fluigent, in partnership with Beonchip, offers a wide range of cell culture chips according to the field of application: to study cell culture under flow, to explore the crosstalk between different 2D and 3D cultures in biomimetic environments, to apply electrochemical gradients to 3D cell cultures, and to study complex culture configurations by joining a culture within a microfluidic channel via a porous membrane.
Optimized cell culture activity: Constant perfusion enables the continuous renewal of nutrients and oxygen to promote cell growth and maintain optimal activity during long-term cell cultures.
Biomechanics: Organ on chip cell culture technology has paved the way for investigating the impact of mechanical strains in cell biology research by reproducing key aspects of an in-vivo cellular microenvironment. Combining microfluidics and microfabrication enables one to reproduce mechanical forces experienced by living tissues at the cell scale.
Passive stimulation induced by shear flow: Liquid flow usually induces shear stress on cells or tissues cultured on the device. This is called shear flow, and the effect is substantial on cell growth, phenotype, and genetic expression.
Active mechanical stimulations originate from the function of the organ. Organs like lungs, muscles and intestines are in active motion. Cells in those organs are mainly subjected to compression and stretching. Organ-on-a-chip cell culture can simulate these environments, allowing for more realistic and detailed results to be obtained.
Biochemical studies: Cells are constantly exposed to biochemical stimulation from the early embryonic stage to adult life. The spatiotemporal regulation of these signals is essential as it determines cell fate, phenotype, metabolic activity as well as pathological behaviors. The fast response and high stability of Fluigent instruments make them the best solution available on the market to reproduce these complex variations in-vitro.
Read our expertise page to know more about the benefits of flow control in cell culture, learn more about our products and those of our partners (Beonchip), and reach out to our team of experts to find a solution that best meets your needs.
Flow control systems and Fluigent’s added value
Implementing perfusion and automated fluid delivery in organ on chip cell culture protocols offers major advantages. Compared to manual pipetting, Automation increases reproducibility, saves time, and improves the level of control in the experiment as all the parameters are tightly regulated (time of delivery, volume, and speed of injection).
Multiple flow control technologies are available for sub-millimeter range fluid management. As demand for microfluidic pumps with higher flow stability, fast response time, versatility, and automation capabilities have increased, pressure controllers have become the device of choice.
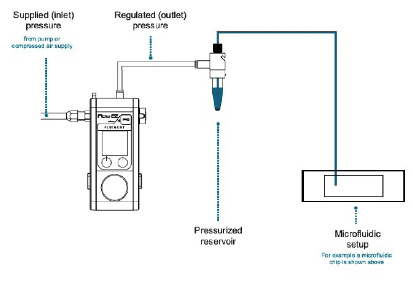
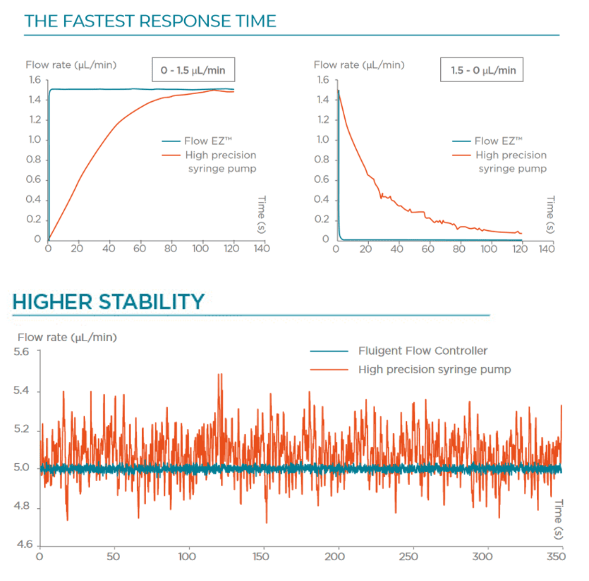
Response time and stability
The working principle of such pumps is to pressurize the sample reservoirs to control the pressure drop between the inlet and the outlet of the microfluidic system. The responsiveness of the generated flow rate depends on the responsiveness of the pressure pump.
Microfluidics in Pharmaceutics
Drug delivery
Microparticles and emulsions are used for a wide variety of pharmaceutical products including intravenous, intramuscular, ocular, or orally delivered compounds. Emulsions are also used as templates for polymer microparticles, lipid nanoparticles, or microcapsules. These are later used for drug delivery, with the emulsion being the active pharmaceutical ingredient (API) itself, or as an adjuvant for co-administration.
Droplet microfluidics technology produces multiunit drug delivery systems with precise dosage control, targeted release, and homogeneous distribution. Conventional methods struggle to achieve monodispersity (<5%) required for efficient drug delivery. Our technology revolutionizes drug delivery, offering improved treatment options and personalized therapies and advancing pharmaceutical development and patient care.
To learn more about droplet microfluidics for pharmaceutical applications, read our white paper about droplet microfluidics.
Disease modeling and characterization
Animal studies are the current standard for evaluating potential treatments, but they often struggle to correlate with human outcomes due to physiological differences and genetic variations. These studies are costly, time-consuming, and raise ethical concerns. Human clinical studies face challenges due to individual diversity and research complexity. Microfluidic models in an organ-on-chip format offer a promising alternative, providing an efficient, cost-effective, and ethical approach for drug discovery and personalized medicine. This technology advances the field of pharmaceutics by enabling the development of predictive methods to evaluate new compounds and therapeutics. It then helps in disease modeling and characterization, enhancing the drug discovery and development process.
To delve deeper into the topic of microfluidics for pharmaceutical applications using organ-on-chip systems, read our OOC white paper.
Related products
Research field

Encapsulation Platform for FACS
Platform for cell encapsulation in DE droplets
See the offer
Microfluidic flow controller
Flow EZ™
See the offer
Microfluidic Double Emulsion Device
RayDrop Double Emulsion
See the offer
Microfluidic Single Emulsion Device
RayDrop Single Emulsion
See the offer
Double Emulsion Generation Pack
Double Emulsion Generation Pack
See the offer
Microfluidic Complex Emulsion Production Platform
Platform for emulsions & droplets
See the offer
PLGA Microparticle Production Pack (Automation Pack)
PLGA Microparticle Production Pack (Automation Pack)
See the offer
Industrial field
Related Ressources
Looking for another market?
From the life sciences to the food industry, many applications require the use of fluids driven at flow rates from nanoliters to milliliters per minute. At such low flows, the success of these applications strongly depends on the level of control and automation of the fluidic operations.
These applications require flow control systems that are adapted for ensuring their success.
In collaboration with the University of Rochester’s Biomedical Engineering department

Testimonial
“While developing a new type of tissue chip that integrates real-time biosensing capabilities, I needed a microfluidic system that could deliver varied samples at precise flow rates. The Aria’s intuitive interface and automation software make it very easy to use for long-term experiments with many steps, and the internal fluidic routing conveniently avoids interfering with our already-complicated optical experimental setup. Also, the flow rate readout and feedback make it easy to control pressure inside our microfluidic devices and prevent bubbles, which is very important for our cell-based experiments. Fluigent was incredibly helpful in supporting us as we introduced their equipment into our lab and continues to provide excellent support when we (rarely) need it.”
John Cognetti, University of Rochester
Why develop a photonic biosensor-integrated tissue chip model?
An alternative to animal studies for drug discovery
Although animal studies are currently the preferred option for evaluating the safety, efficacy, pharmacokinetics, and toxicity of potential treatments, their results are often difficult to correlate with human outcomes due to physiological differences and genetic variations between species. Additionally, animal studies can be costly, require a large workforce, and raise ethical concerns due to the need for a significant number of animals. Human clinical studies also face challenges due to the heterogeneity of individuals and the complexity of the studies. Therefore, microfluidic models in a chip format may offer an efficient alternative to animal studies, providing a reliable, cost-effective, and ethical approach for drug discovery.
Advancements in In Vitro Models: Tissue Chips Mimicking Human Organs and Tissues
In vitro models are an alternative to animal models that can overcome the problem of heterogeneity between humans and animals and can also reduce costs and avoid ethical issues. However, previous in vitro models were limited in complexity and could not reflect physiological relevance of in vivo tissues or organs. To address this issue, microfluidic technologies, such as the tissue chip platform developed by the Biomedical Engineering department at the University of Rochester, were developed. This technology incorporates a microfluidic delivery system capable of mimicking the complexityof human organs and tissues, including 3D architectures.[1] These chips provide a physiologically relevant level of tissue organization, geometry, soluble gradients, and mechanical stimulation in an accurate and regulated manner. [2] This technology represents a significant advancement in drug discovery and development, as it allows for the development of more predictive methods for assessing the toxicity and efficacy of new compounds and therapeutics, as well as disease modeling and characterization. [1] Moreover, the integration of sensors into these microfluidic chips enables high throughput analysis.
Benefits of using a photonic biosensor integrated into the chip
The primary objective of this study is the real-time sensing of specific biomolecules. Detecting cytokine secretion by epithelial cells in real-time is a crucial goal for tissue chip technologies, as cytokines play a vital role in various inflammatory diseases such as lung infections and can act as useful biomarkers. Unfortunately, few techniques have been developed to determine the cellular response to a stimulus in real time, and the few that exist may utilize fluorescent tags that can disturb the system or lack specificity in their biosensing capabilities. [4] As a result, researchers from the Department of Biomedical Engineering at the University of Rochester have taken up this challenge and developed a new platform based on tissue chip technology. The purpose of this tissue chip platform is to measure the secretion kinetics and concentrations of analytes in close proximity to the lung epithelial cells, which will aid in better understanding the disease dynamics. To maximize the sensing performance, a photonic biosensor has been integrated into the chip at extreme proximity to the cells, which represents an innovative, label-free, and highly sensitive solution.
Using the ARIA in the tissue chip platform for real-time sensing of inflammatory markers
Description of the tissue chip platform
The University of Rochester has developed a new, innovative platform with the goal of improving existing tissue chip models and creating a device capable of sensing the secretions related to the inflammation of lung epithelial cells in real time. This platform consists of various components listed below:
- A microfluidic tissue chip: This microfluidic device includes two microfluidic channels separated by a thin nanoporous membrane. The upper channel is used for sample supply, while the lower channel is sealed and in direct contact with the biosensor, which collects the inflammatory markers secreted by the cells. The lowest layer of the chip is equipped with a photonic biosensor that can capture certain resonance wavelengths that are modified by the refractive index of entities in close proximity, thanks to 14 rings functionalized according to the targeted analyte. Thus, in the presence of cytokines around the rings, a shift in the resonance wavelengths of the rings towards the red end of the spectrum may be observed in a manner consistent enough to be quantified according to the concentration of cytokine in the medium. The membrane is made of a nanoporous silicon nitride membrane (NPN), optically clear, composed of four slots and serves as a substrate for the cell culture. The cells will be able to adhere to it and create tight junctions in the form of a monolayer network.
- Fluid delivery: To supply the tissue chip device continuously during the experiment, our automated sequential injection system, the ARIA, is used upstream of the chip. It is controlled by a pressure-driven fluid pump and can be pre-loaded with up to 10 different samples. A switch function allows for seamless transition between samples, and a bubble trap prevents air from disrupting the cells in the device.
- Optical configuration: An optical set-up is also used to capture the resonance shifts transmitted by the sensors in real time. This includes a tunable laser source, a fiber array aligned to the chip, and visible IR cameras.
- Data analysis: Finally, a custom python program is used to compute the resonance shifts transmitted by the sensors and captured by the optical set-up, allowing the experimenter to access this data via an interface. The interface allows users to select the peaks and categorize them according to the nature of the sensor ring (control or capture) and thus the relative shift is calculated depending only on the specific capture of the analytes.
The tissue chip platform is placed in an aluminum stage fitted with a thermoelectric coupler and a thermistor to maintain the device at 37°C.
Course of the experiment
Human bronchial epithelial cells (16HBE line) are used in the tissue chip model due to their ability to form tight monolayers in vitro, attributed to the presence of a robust network of tight junctions. These cells are also known to secrete cytokines in response to stimulation with bacterial endotoxin lipopolysaccharide (LPS). A complete or nearly complete monolayer of cells is seeded into the device, covering the four membrane slots. The Aria reservoirs are preloaded with the necessary solutions (media, LPS, controls) and the platform is maintained at 37°C. During sensing experiments, the two channels are disconnected from each other. The bottom channel is sealed with binder clips to capture analytes produced during passive cell diffusion on the photonic biosensor, while the inlet of the top channel is connected to the outlet tubing of ARIA. The medium is diffused at a continuous rate of 30 µL/min through the top channel during 30min, followed by the LPS at 100 ng/mL for two hours. By aligning the chip to the fiber array, spectral measurement can begin immediately, and data is acquired in real-time.
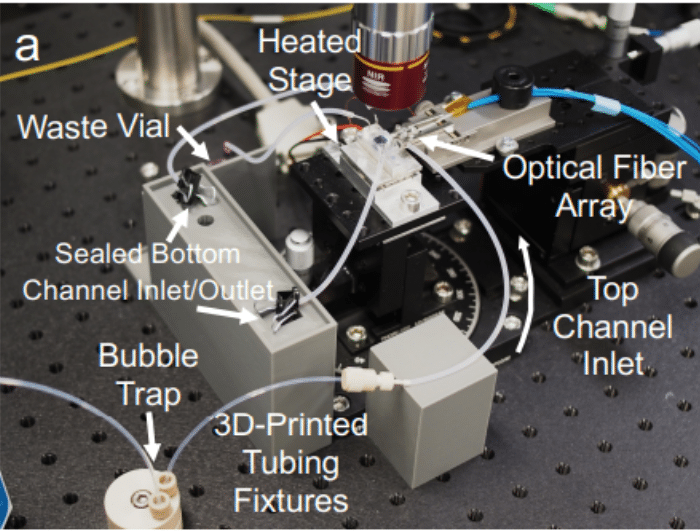

Figure 3: Set-up of the platform.
Partial results
Control tests
Initially, a single channel microfluidic device was used to calibrate the sensors with four analytes. The pro-inflammatory cytokines IL-1β, IL-6, and IL-8 were tested, along with the C-reactive protein (CRP), which acted as a negative control for cell stimulation (not secreted by the 16HBE), and a positive control for the sensor (120kDa pentamer easily detected by the photonic biosensor; added exogenously at the end of the experiment). The four tests confirmed that the relative shift in pm is directly proportional to the analyte concentration. Next, a second test was conducted to ensure that the 16HBE cells behaved as expected and formed tight junctions. This was proven using the junction marker ZO-1 after four days of cell culture in the device. Finally, a control was performed to prove that analytes originating from the membrane can indeed reach the membrane in a short time. The assay was conducted on the analytes IL-1β, IL-6, and IL-8 in a cell-free chip, and relatively similar shifts were observed after 5-10 minutes after the entry of the different analytes into the top channel. This validated the possibility of acquiring time-resolved data of the cellular response to a stimulus added to the top channel.
The tests confirmed the proper functioning of the different key components of the tissue chip device. The next step involves testing the platform under experimental conditions.
Sensing of cellular secretion partial results
The media flowed via the ARIA for 30 minutes on the complete platform described above, followed by the injection of LPS at 100ng/mL for two hours. One hour after the injection of LPS, a sensor response was observed, which resulted in an increase in the relative shift for interleukin IL-1β and IL-6 secreted by the lung epithelial cells. Equilibrium was reached between 90-120 minutes. This demonstrates that interleukins can diffuse to the biosensor located at the bottom of the tissue chip. The response was shown to be specific to the cytokines secreted by HBEs by analyzing the curve in Figure 4c, which shows the lack of variation of the relative shift as a function of the CRP concentration. An additional test was performed using a membrane without cells on some of its slots, revealing that the shift is less strong for the sensors farther away from cells.
In summary, these experiments have demonstrated that the photonic tissue chip platform can detect physiologically relevant concentrations of analyte and report on the quantity of cytokines secreted by inflamed epithelial cells in real time following LPS stimulation. Additional tests have also shown that these sensors can provide data with both spatial and temporal resolutions.
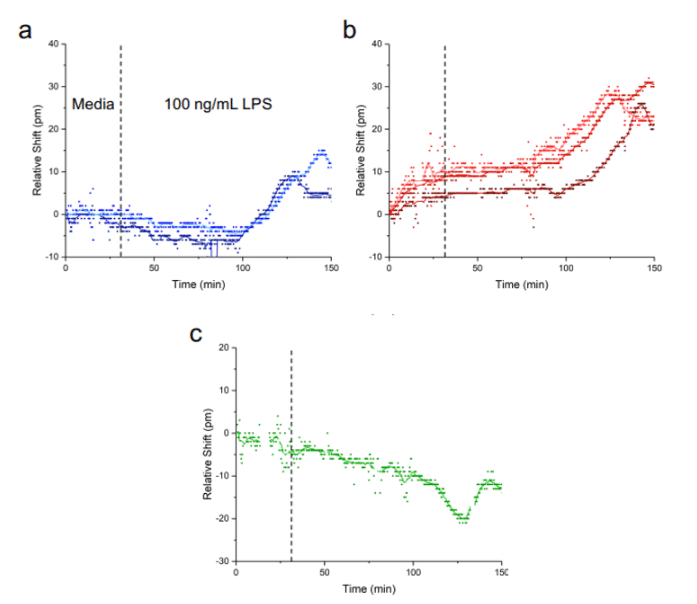
References
[1] Low LA, Tagle DA. Tissue chips – innovative tools for drug development and disease modeling. Lab Chip. 2017;17(18):3026-3036. doi:10.1039/C7LC00462A
[2] Donoghue L, Nguyen KT, Graham C, Sethu P. Tissue Chips and Microphysiological Systems for Disease Modeling and Drug Testing. Micromachines 2021, Vol 12, Page 139. 2021;12(2):139. doi:10.3390/MI12020139
[3] Microfluidic white paper – A guide to Organs-on-Chips technology, Fluigent
[4] Morales AW, Zhang YS, Aleman J, et al. Label-free detection of protein molecules secreted from an organ-on-a-chip model for drug toxicity assays. In: Frontiers in Biological Detection: From Nanosensors to Systems VIII. Vol 9725. SPIE; 2016:972508. doi:10.1117/12.2212971