Highly monodisperse droplets
PDMS Drop-seq chip for Drop-seq experiments
[ODROPSEQC]Drop-seq chip
Our Drop-seq chip is a PDMS Chip containing 22 operational designs including a Silane hydrophobic coating.
Drop-seq chip advantages:
-
- High throughput results in a short period of time
- Minimize consumption of expensive samples
- Prepare a single-cell suspension from a tissue
- Analyzes sequences of single cells in a highly parallel manner
- Unique molecular and cell barcodes enable cell- and gene-specific identification of mRNA strands
- RT with template-switching PCR produces high-yield reads from single cells
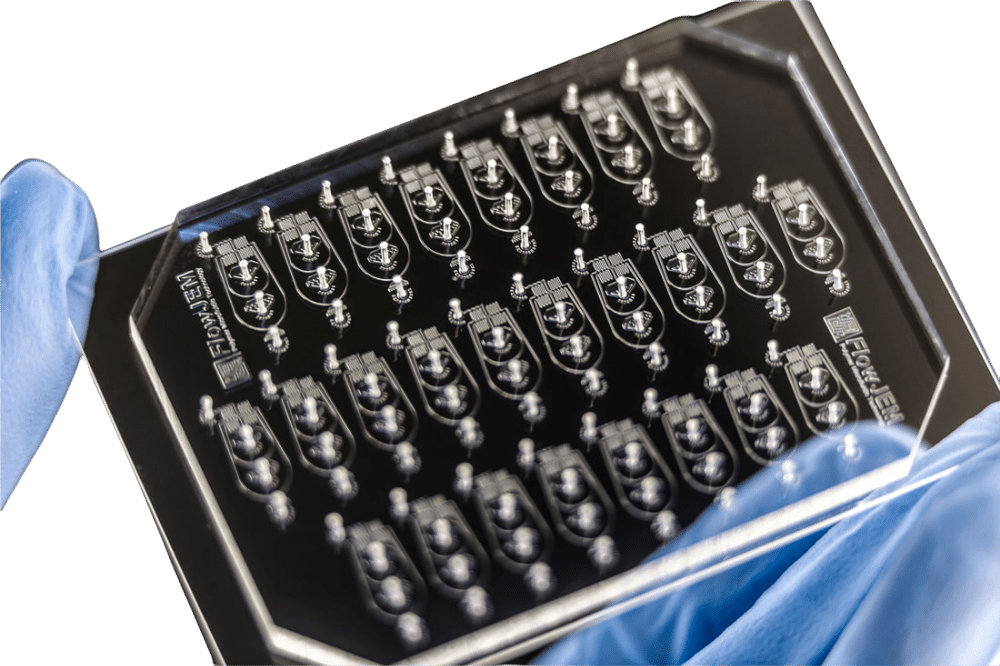
- Monodispersity
- Robust
Durable over a wide range of parameters
- Efficient
Efficient production of transcript libraries
Single-cell RNA-sequencing (scRNA-seq) has revolutionized biomedical research by enabling the in-depth analysis of cell-to-cell heterogeneity of tissues with unprecedented resolution. Among several existing droplet-based approaches, the Drop-seq method, based on the generation of water-in-oil emulsions, has emerged as one of the most widely used systems.
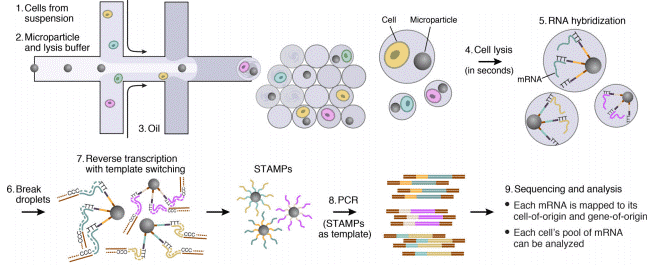
The single-cell sequencing device is a low-cost, high-throughput platform to profile thousands of cells by encapsulating them into individual droplets. Uniquely barcoded mRNA capture microparticles and cells are coconfined through a microfluidic device within the droplets where they undergo cell lysis and RNA hybridization. After breaking the droplets and pooling the hybridized particles, reverse transcription, PCR, and sequencing in single reactions allow to generate data from thousands of single-cell transcriptomes while maintaining information on the cellular origin of each transcript. More details about the mechanism and application of Drop-seq.
What is PDMS and why use it for Drop-seq chip?
PDMS is used to fabricate microfluidic devices (single layer and bilayer) and micro-imprint stamps. Two different types are commonly used by researchers for these applications: PDMS RTV-615 and PDMS Sylgard 184.
- It is deformable, which allows the integration of microfluidic valves using the deformation of PDMS micro-channels, its use to detect very low forces (biomechanics interactions from cells)…
- It is easy to mold. The PDMS can mold structures at high resolutions. Also, PDMS bonds tightly to glass or another PDMS layer with a simple plasma treatment.
- It is inexpensive compared to other materials.
- It is generally biocompatible.
Features of Drop-seq chip
Dedicated to Drop-seq
Each Droplet Generation Device is based on the design recommended in the latest McCarroll lab Drop-seq protocol, ensuring the best chances of success.
More than 22 experiments per chip
22 Droplet Generation Devices per Chip Provides value for money in a chip that lasts. When the life of one device is depleted, simply move on to the next one. Besides, our drop-seq chip rapidly creates libraries ready for high throughput sequencing.
Efficient production of transcript libraries
Superior design promotes optimal mixing of component fluids, thereby minimizing bead shearing or premature lysis of cells and mRNA release.
Libraries from each cell are uniquely bar coded and they can practically barcode over a million cells per experiment.
The library generation and bar coding workflow is fast, simple, rugged and robust.
Drop-seq Applications
Identification of cell subpopulations
One of the main applications of Drop-seq chip is in the study of cell populations and the identification of cell subpopulations. By analyzing the transcriptomic profiles of individual cells, researchers can identify cells with similar gene expression patterns and classify them into subpopulations. This can be useful in understanding the diversity and function of cells in complex tissues, such as the brain or the immune system.
Cell differenciation analysis
Another application of Drop-seq is in the study of developmental processes. By isolating and sequencing individual cells at different stages of development, researchers can understand how gene expression changes over time and how cells differentiate into different cell types. This can provide insight into how organisms develop and how different cell types are formed.
Cancer research
The drop-seq chip has also been used in the field of cancer research, where it can be used to identify subpopulations of cancer cells with distinct gene expression patterns. This can help to understand the heterogeneity of cancer and to identify potential targets for therapy.
Furthermore, Drop-seq has been applied in several other fields such as:
- Immune function: Drop-seq has been used to study the immune system by isolating and sequencing individual immune cells. This can help to understand the diversity and function of immune cells, such as T cells and B cells, and how they respond to different pathogens.
- Stem cells: the single-cell sequencing device has been used to study stem cells and their potential to differentiate into different cell types. By isolating and sequencing individual stem cells, researchers can understand how gene expression changes as stem cells differentiate and how this process is regulated.
- Novel cell types: The drop-seq chip has been used to identify novel cell types in complex tissues, such as the brain. By analyzing the transcriptomic profiles of individual cells, researchers can identify cells with distinct gene expression patterns that may represent novel cell types.
- Microbiome: Drop-seq has been used to study the gut microbiome by isolating and sequencing individual gut bacteria. This can help to understand the diversity and function of gut bacteria and how they interact with host cells.
Overall, the single-cell sequencing device is a powerful tool for studying cell populations and understanding the diversity and function of cells at a single-cell resolution. It has a wide range of applications in many different fields, from understanding the function of cells in complex biological systems to identifying novel cell types and drug resistance.
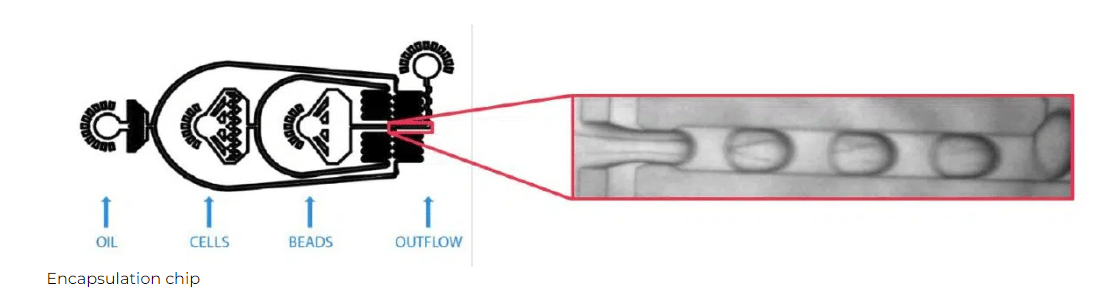
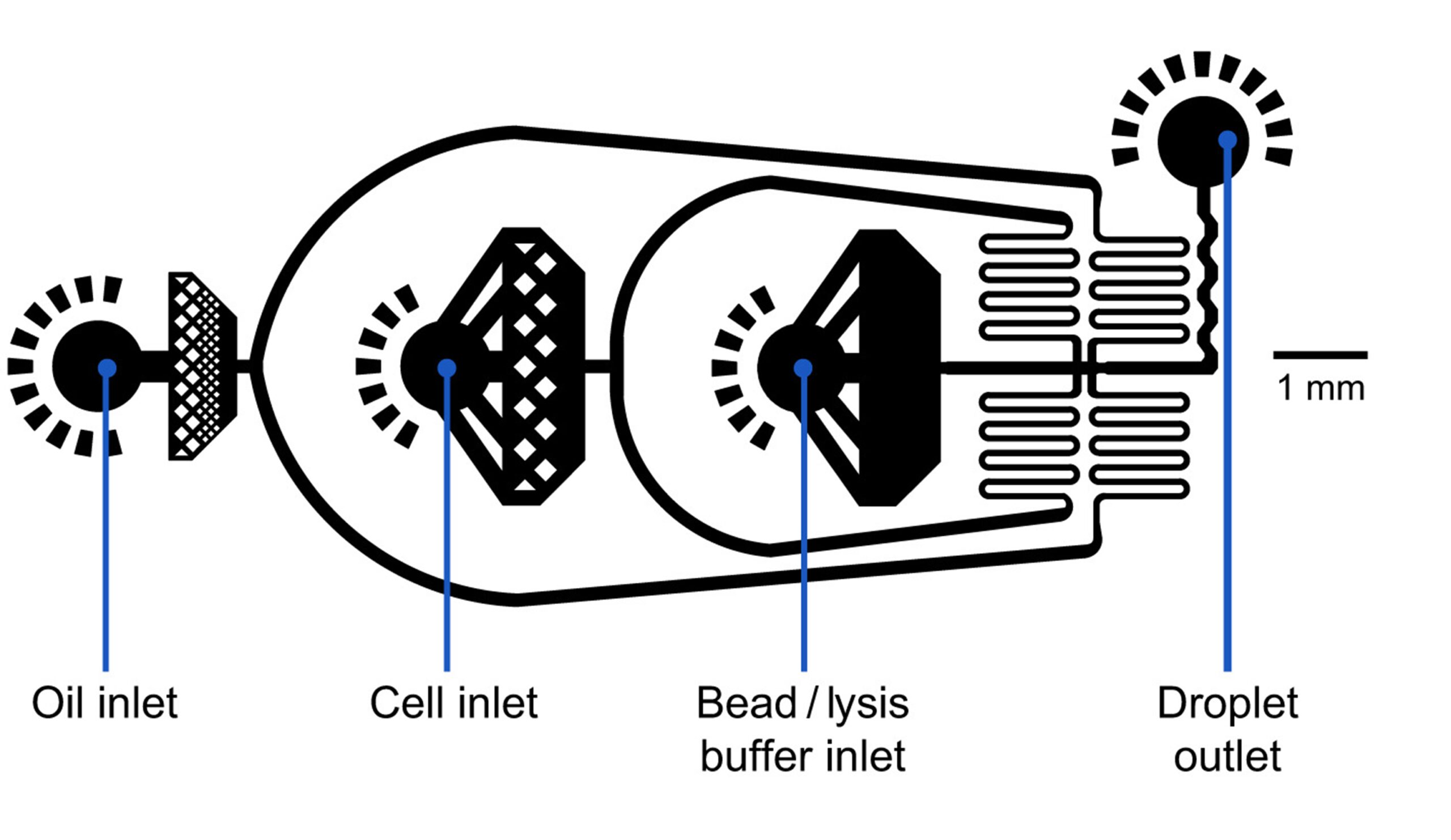
Specifications
Chip Design
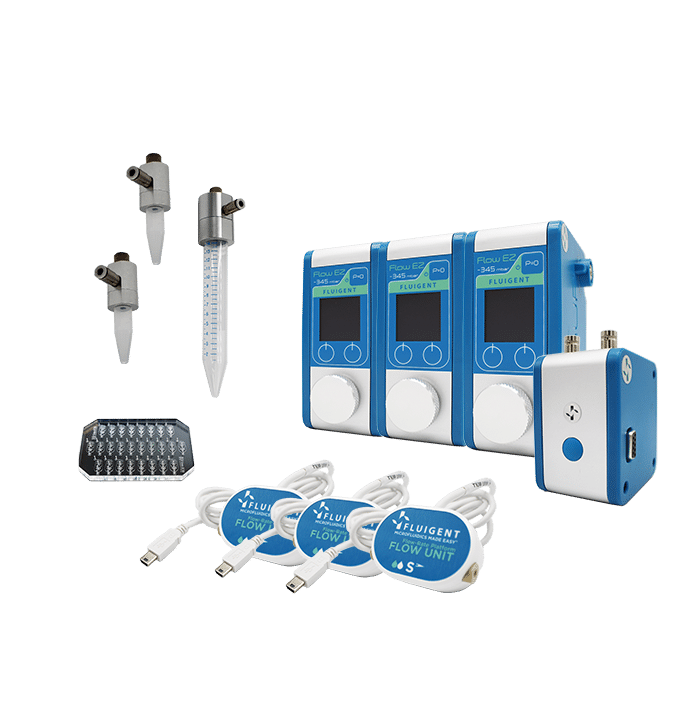
Drop seq package setup
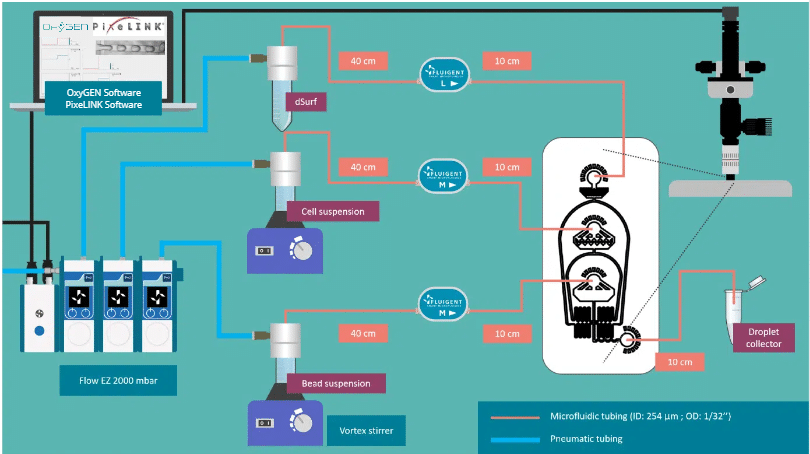
Expertise & resources
-
Fluigent Products Datasheets Drop-seq chip datasheet Download
-
Microfluidic Application Notes Droplet Sequencing: Drop-Seq method Read more
-
Fluigent products manual McCarroll Drop-seq protocol Download
-
Fluigent products manual Macosko Drop-seq article Download
-
Expert Reviews: Basics of Microfluidics Microfluidic Droplet Production Method Read more
-
Expert Reviews: Basics of Microfluidics Flow control for droplet generation using syringe pumps and pressure-based flow controllers Read more





