Microfluidics for Organ-on-chip Cell culture
The science of flow control for organ-on-a-chip cell culture & its applications
- In-vitro model
- Better mimicked and controlled microenvironment
- Cost-effective
- Scaling possible
Organ-on-a-chip (OOAC) is the concept of mimicking the organ-level function of human physiology or disease using cells inside a microfluidic chip. Microfluidics provides the unique ability to control the cellular microenvironment with high spatiotemporal precision and to present cells with mechanical and biochemical signals in a more physiologically relevant context. The ability to manipulate micro-liter volumes of liquids has made these models a platform where scaling and dynamic crosstalk between cells can be achieved.
Microfluidic chips can now use geometries and structures to permit the use of physiological length scales, concentration gradients, and the mechanical forces generated by fluid flow to mimic the in vivo microenvironment experienced by cells. These biomimetic platforms overcome many of the drawbacks encountered with conventional tissue culture models.
Applications of OOAC cell culture
Therapy development
Organ-on-chip cell culture platforms have proven potential in providing tremendous flexibility and robustness in drug screening and development by employing engineering techniques and materials. More importantly, there is a clear upward trend in studies that utilize human-induced pluripotent stem cells (hiPSC) to develop personalized tissue or organ models. For this purpose, the use of cell culture chips allows users to study complex culture configurations by joining a culture well with a microfluidic channel via a porous membrane. This is the optimal device for Air Liquid Interface (ALI) culture, endothelium/epithelium barrier and crosstalk studies.

Drug discovery
The development of emerging in-vitro tissue culture platforms can be useful for predicting the human response to new compounds. Recently, several in-vitro tissue-like microsystems, also known as “organ-on-a-chip studies”, have emerged to provide new tools for better evaluating the effects of various chemicals on human tissue.
Organ-on-chip cell culture models can therefore be used for accurate prediction and mechanistic investigation of dose-limiting human toxicities of prospective drugs, as well as for the exploration of new therapeutic approaches to mitigate the observed toxic effects. In the drug discovery pipeline, predictions made by these models could inform and facilitate early efforts to identify, modify and optimize lead compounds, thereby developing safer drugs with an increased likelihood of success in clinical trials.
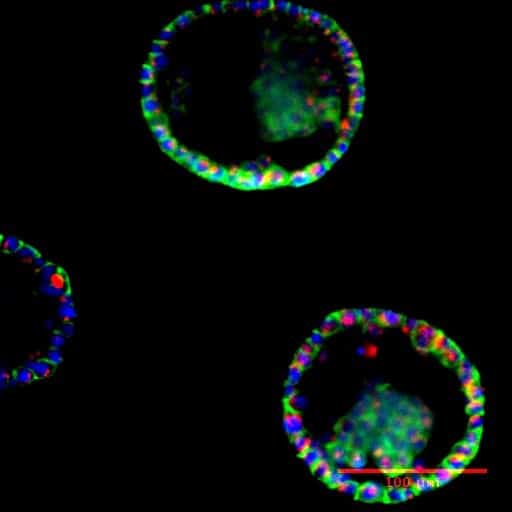
Regenerative medicine
The development of emerging in-vitro tissue culture platforms can be useful for predicting the human response to new compounds. Recently, several in-vitro tissue-like microsystems, also known as ‘organ-on-a-chip studies’, have emerged to provide new tools for better evaluating the effects of various chemicals on human tissue.
Organ-on-chip cell culture models can therefore be used for accurate prediction and mechanistic investigation of dose-limiting human toxicities of prospective drugs, as well as for the exploration of new therapeutic approaches to mitigate the observed toxic effects. In the drug discovery pipeline, predictions made by these models could inform and facilitate early efforts to identify, modify and optimize lead compounds, thereby developing safer drugs with an increased likelihood of success in clinical trials.
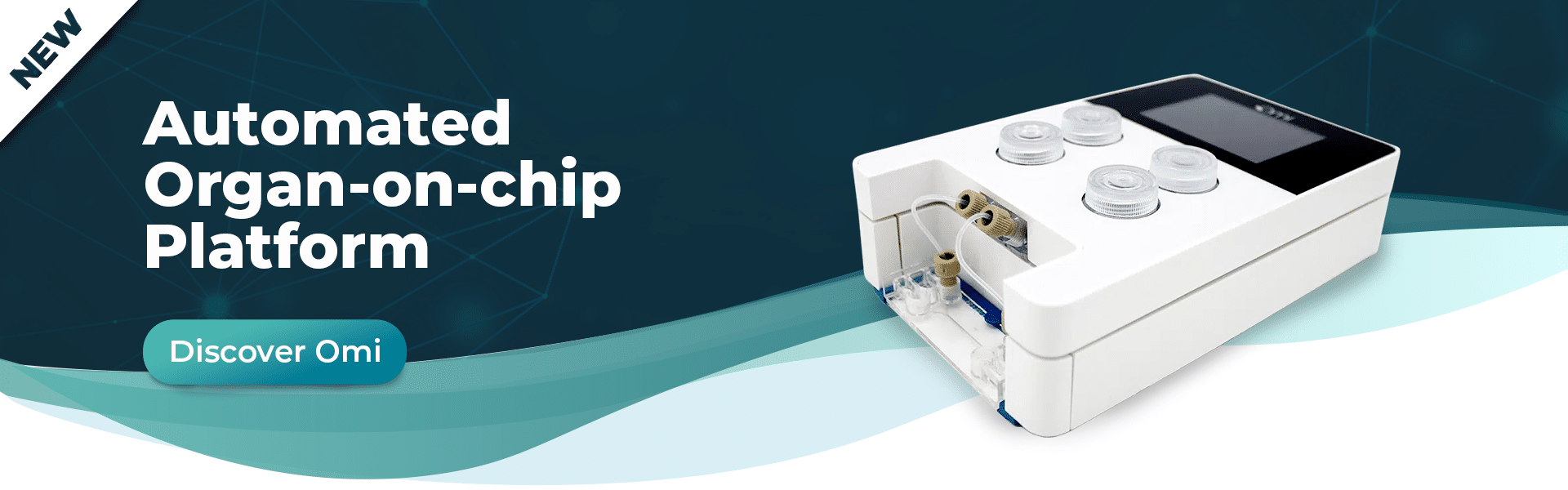
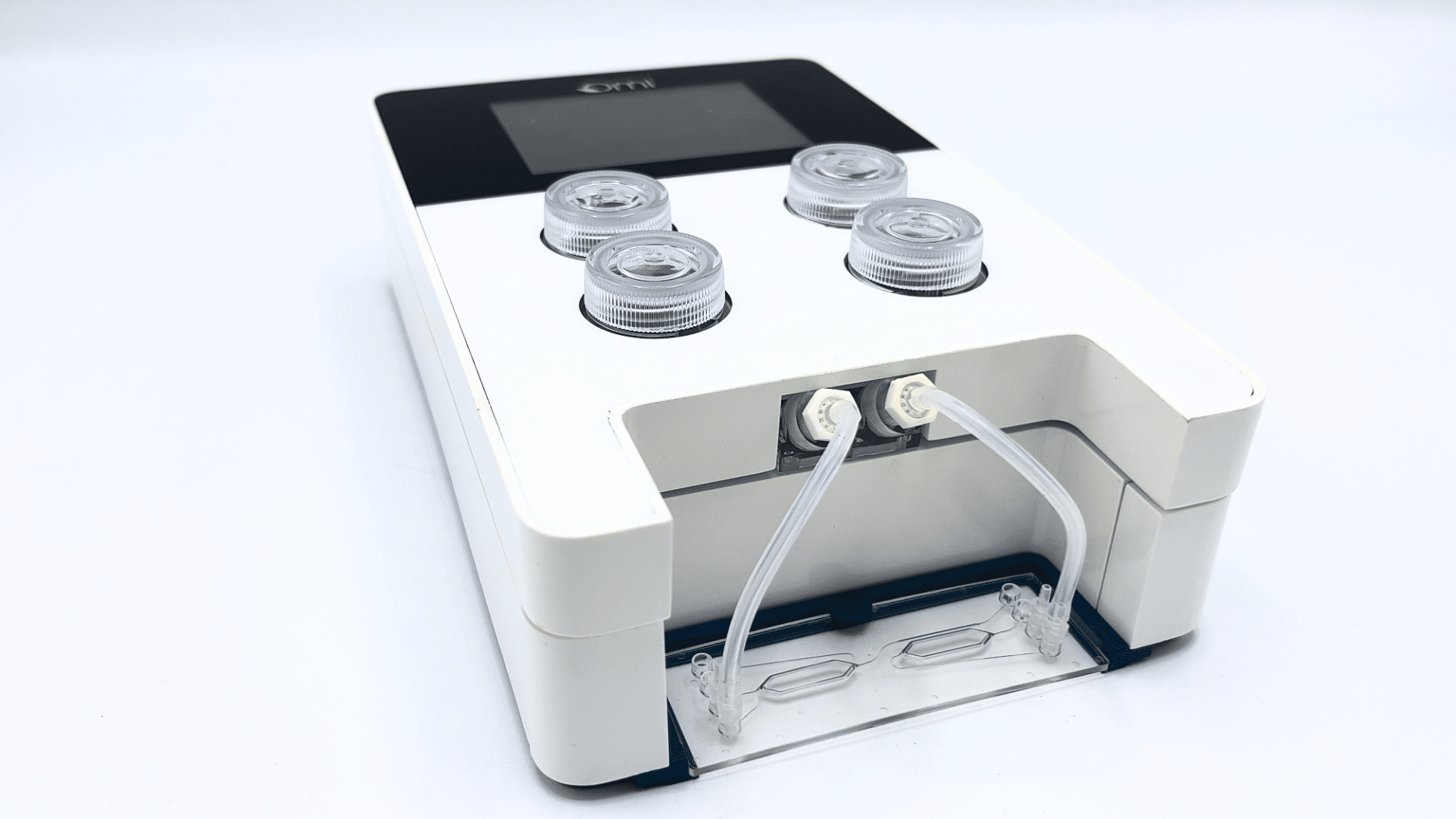
Omi, the new automated organ-on-chip platform
Discover Omi, the new automated platform for OOAC applications developed by Fluigent. Equipped with state-of-the-art technologies, this platform will enable you to carry out any perfusion protocol. It has the ability to customize and automate any protocol, including simple perfusion, recirculation, sampling and injection. It meets the needs of beginners in organ-on-chip cell culture research and advanced organ-on-chip researchers looking for automation and reproducibility.
This versatile, automated organ-on-a-chip platform can perform long-term OOAC cell cultures under flow to control shear stress conditions. Its two-hour battery life and WiFi connectivity ensure easy, intuitive control. It can be easily transported from incubator to microscope for live cell imaging while maintaining cell perfusion under battery power. You can also monitor your experiment from anywhere using the Omi app.

Towards the next generation of organ-on-a-chip cell culture platforms
Fluigent and Beonchip are partnering to offer a complete solution for organ-on-chip cell culture.
This webinar will first introduce Beonchip, their chips and the numerous applications that can be performed with them.
The second part will focus on flow control systems. Although often considered secondary, flow control is as important as chip design, as cells are highly sensitive to mechanical forces. Results demonstrate how cells are affected by peristaltic pumps compared to pressure control systems.
Organ-on-chip research is an emerging field that offers substantial benefits compared to conventional cell culture. In many labs, considerable effort is put into choosing the right chip design, but the impact of flow control is still undetermined.
It is our intent to create awareness of the importance of flow and its effects on studies.
Read our expertise page to learn more about the benefits of flow control in cell culture and about our products and those of our partners (Beonchip), and to reach out to our team of experts to find the solution that best meets your needs.
Resources
-
Microfluidic Application Notes Long-term fluid recirculation system for Organ-on-a-Chip applications Read more
-
Microfluidics case studies CNRS/UTC: study of a liver-on-a-chip model Read more
-
Expert Reviews: Basics of Microfluidics Why is a controlled shear-stress a key parameter of your microfluidic experiments? Read more
-
Microfluidic Application Notes Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions Read more
-
Microfluidic Application Notes Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications Read more
-
Microfluidic Application Notes Assess Cell Proliferation Using Pressure as a Tool Read more
-
Microfluidic Application Notes Cartilage-on-a-chip, an example of complex mechanical stimulation using Fluigent’s technology Read more
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Prostate Organoid Culture in Microbeads Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
-
Expert Reviews: Basics of Microfluidics MYOCHIP | H2020 European project Read more
-
Expert Reviews: Basics of Microfluidics Application of microfluidic chip technology Read more
Importance of fluid handling for organ-on-a-chip cell culture
In many labs, considerable effort is put into choosing the right chip design, but the impact of flow control is still undetermined. It is our intent to create awareness of the importance of flow and its effects on one’s studies.
Fluigent, in partnership with Beonchip, offers a wide range of cell culture chips according to the field of application: to study cell culture under flow, to explore the crosstalk between different 2D and 3D cultures in biomimetic environments, to apply electrochemical gradients to 3D cell cultures, and to study complex culture configurations by joining a culture within a microfluidic channel via a porous membrane.
Optimized cell culture activity: Constant perfusion enables the continuous renewal of nutrients and oxygen to promote cell growth and maintain optimal activity during long-term cell cultures.
Biomechanics: Organ on chip cell culture technology has paved the way for investigating the impact of mechanical strains in cell biology research by reproducing key aspects of an in-vivo cellular microenvironment. Combining microfluidics and microfabrication enables one to reproduce mechanical forces experienced by living tissues at the cell scale.
Passive stimulation induced by shear flow: Liquid flow usually induces shear stress on cells or tissues cultured on the device. This is called shear flow, and the effect is substantial on cell growth, phenotype, and genetic expression.
Active mechanical stimulations originate from the function of the organ. Organs like lungs, muscles and intestines are in active motion. Cells in those organs are mainly subjected to compression and stretching. Organ-on-a-chip cell culture can simulate these environments, allowing for more realistic and detailed results to be obtained.
Biochemical studies: Cells are constantly exposed to biochemical stimulation from the early embryonic stage to adult life. The spatiotemporal regulation of these signals is essential as it determines cell fate, phenotype, metabolic activity as well as pathological behaviors. The fast response and high stability of Fluigent instruments make them the best solution available on the market to reproduce these complex variations in-vitro.
Read our expertise page to know more about the benefits of flow control in cell culture, learn more about our products and those of our partners (Beonchip), and reach out to our team of experts to find a solution that best meets your needs.
Flow control systems and Fluigent’s added value
Implementing perfusion and automated fluid delivery in organ on chip cell culture protocols offers major advantages. Compared to manual pipetting, Automation increases reproducibility, saves time, and improves the level of control in the experiment as all the parameters are tightly regulated (time of delivery, volume, and speed of injection).
Multiple flow control technologies are available for sub-millimeter range fluid management. As demand for microfluidic pumps with higher flow stability, fast response time, versatility, and automation capabilities have increased, pressure controllers have become the device of choice.
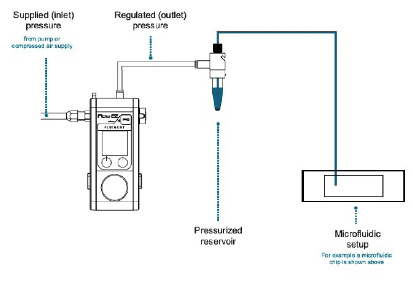
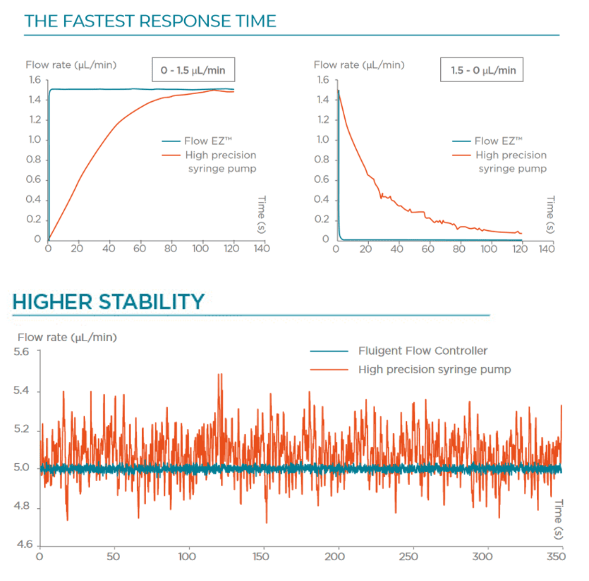
Response time and stability
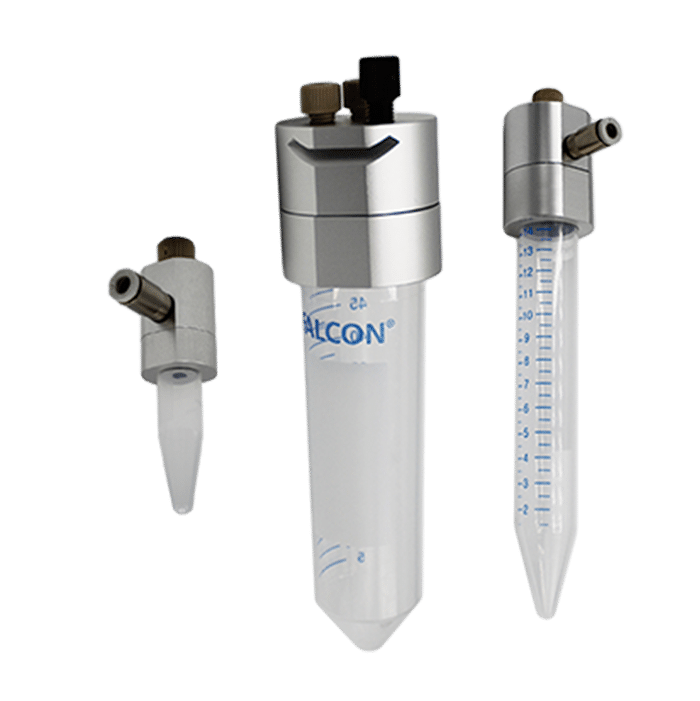
The working principle of such pumps is to pressurize the sample reservoirs to control the pressure drop between the inlet and the outlet of the microfluidic system. The responsiveness of the generated flow rate depends on the responsiveness of the pressure pump.
ALI cell culture chip features
Easy to use
Thanks to its design, the Be-transflow chip can be used for a wide range of applications and with any type of optical microscopy. In addition, its slide format has been chosen to ensure easy and efficient handling, particularly when used under a microscope.
Easy to connect
The cell culture chip is compatible with all Fluigent pressure-based flow controllers. In fact, it is possible to apply a flowrate connecting any Fluigent’s flow controller with the chip using patented inlet/outlets that avoid the entrance of bubbles to the channel.
No unspecific absorption
Unlike in other PDMS devices, the air liquid interface cell culture chip is made of lipophobic thermoplastic materials. Due to that feature, it does not present unspecific drug absorption issues. This enables a wide range of applications, including for instance immunohistochemistry with fluorescent detection.
Cell recovery
The air liquid interface cell cultures used in the Beonchip device can be easily recovered for further experimentation. Indeed, with its channel outlet, the culture medium can then be efficiently isolated and studied, while limiting losses and contamination.
Easy to implement
The wells of the chip are positioned in standard positions of a 96 well plate to facilitate its use, in particular in automated microscopes. In addition to that, their volume is identical to the one of those wells in order to ease the transition to the transflow device, while avoiding loss of cells or media.
Related applications
Air liquid interface culture models
Perform ALI experiments on a 2D or 3D culture with automatic culture medium replacement for systems such as epithelial cultures, toxicity tests, absorption test and much more.
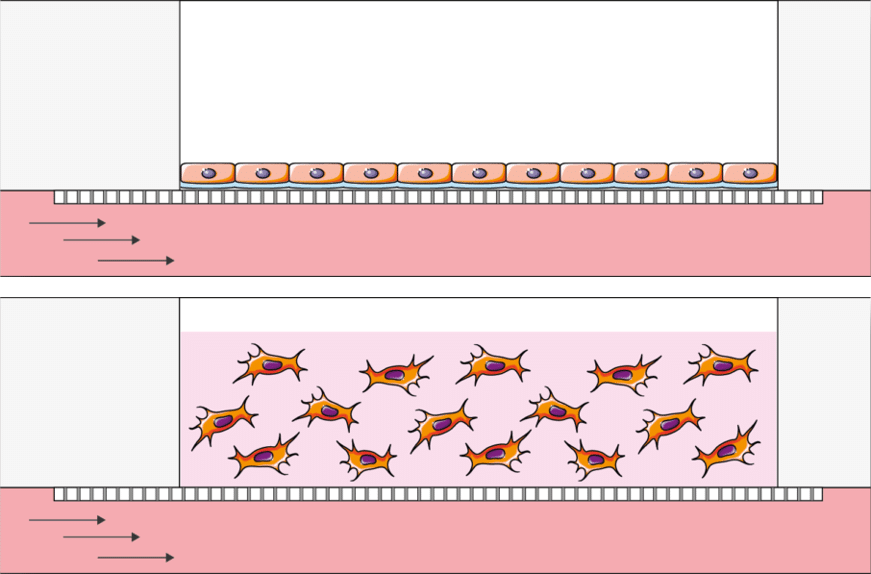
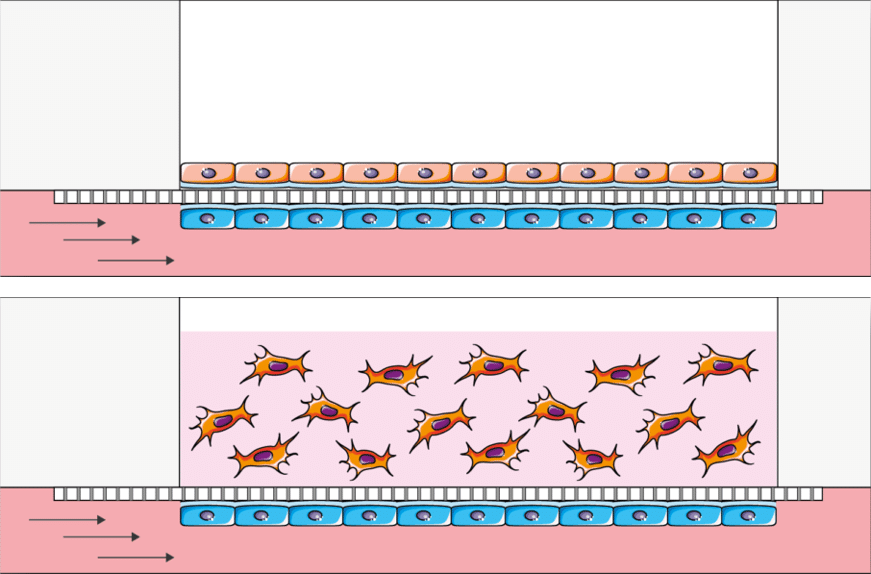
Create the perfect endothelium endothelium-epithelium barrier: Use the upper well for the 2D or 3D epithelial culture and seed the endothelial cells in the perfusable channel underneath. Most common applications are Blood Brain Barrieer (BBB), gut, skin, lung and many more.
Crosstalk studies (prototype): Explore the cross-talk of two cultures in an automated and simple way using the BE-Transflow option that connects two wells in the same channel. Most common application are indirect toxicity studies. This air liquid interface cell culture chip is a prototype and is available at a higher price than the normal BE-Transflow.
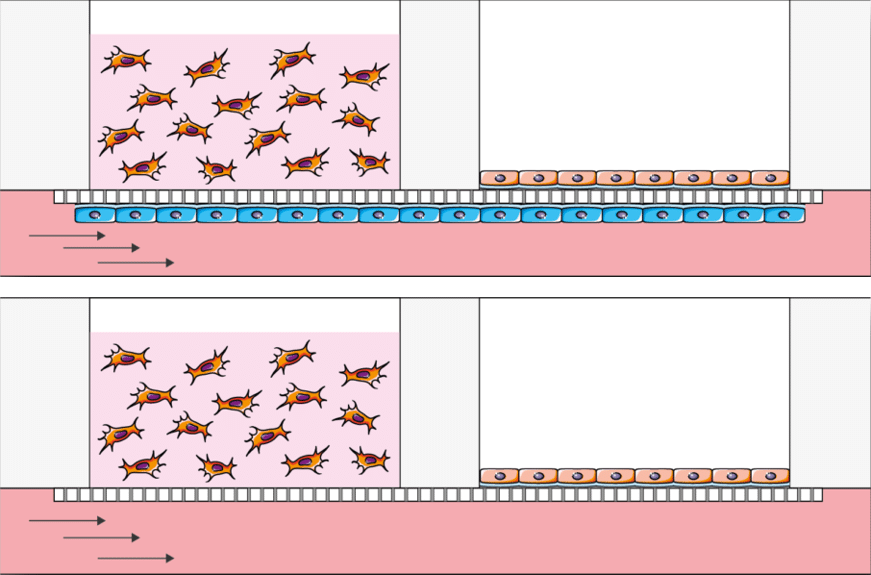
Be-Transflow cell culture chip Applications
BE-Transflow has been designed with ALI in mind. Its an ideal chip for ALI cell culture and coculture research (skin, cornea, gut, lung). However, its simplicity to use makes it also a very useful tool for any coculture research and crosstalk experiments.
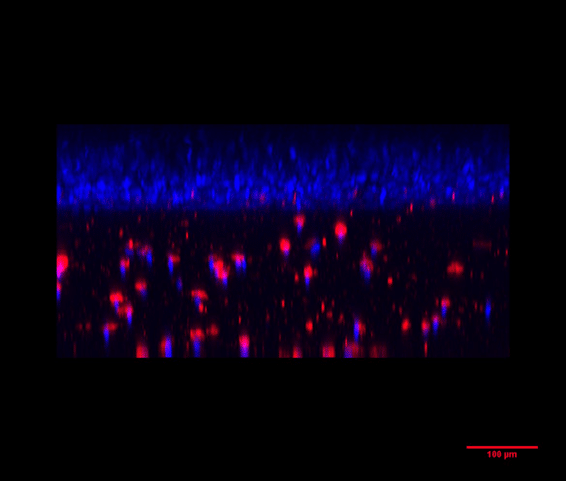
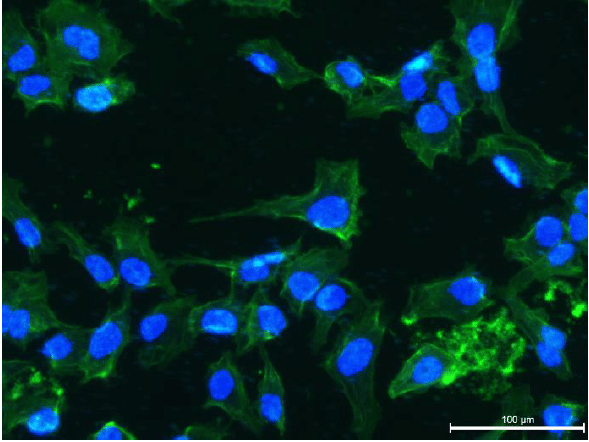
Epidermis/dermis coculture: Recreation of the epidermis and dermis in the well of the BE-Transflow device. In the upper part of the image and marked in blue are the nuclei of the keratinocytes (HEKa) forming the epidermis. The HEKa cells were cultured for 21 days in the ALI air liquid interface to create a mature epidermis. Underneath the epidermis we can see the fibroblasts in red embedded in a collagen matrix, mimicking the dermis. The scale bar marks 100 µm.

Bone on chip: BEOnchip participates in the Eurostars project BONAFIDE. This project is devoted to the production of a human bone-on-chip platform that will simulate bone growth, resorption and remodeling for future evaluation of (anti-osteoporosis) drugs and biomaterials (e.g. bone cements).
The air liquid interface cell culture well represents the catabolic side (bone resorption) where the osteoclasts rest in contact with a bonelike scaffold. The lower channel represents the anabolic side of the bone model. This channel is seeded with osteocytes and osteoblast that form a 2D tissue.
This tissue is mechanically stimulated by the controlled laminar flux applied in the channel, mimicking the physiological environment of the bone. The bone remodeling process is automatically analyzed using image analysis software and biomarker analysis.
Specifications
PERFORMANCE
| Height | Width | Length | Total volume | |
| Channel | 375 µm | 1.5 mm | 45 mm | 44 µL |
| Well | 6 mm | 5.7 mm | 5.7 mm | 195 µL |
| Inlet/outlet | 7 mm | UNF 1/4″ – 28 | UNF 1/4″ – 28 | 130 µL |
| Medium reservoir | 7 mm | 3.6 mm | 8.8 mm | 185 µL |
| Membrane pore size | 1 µm | 1 µm | 1 µm | 1 µm |
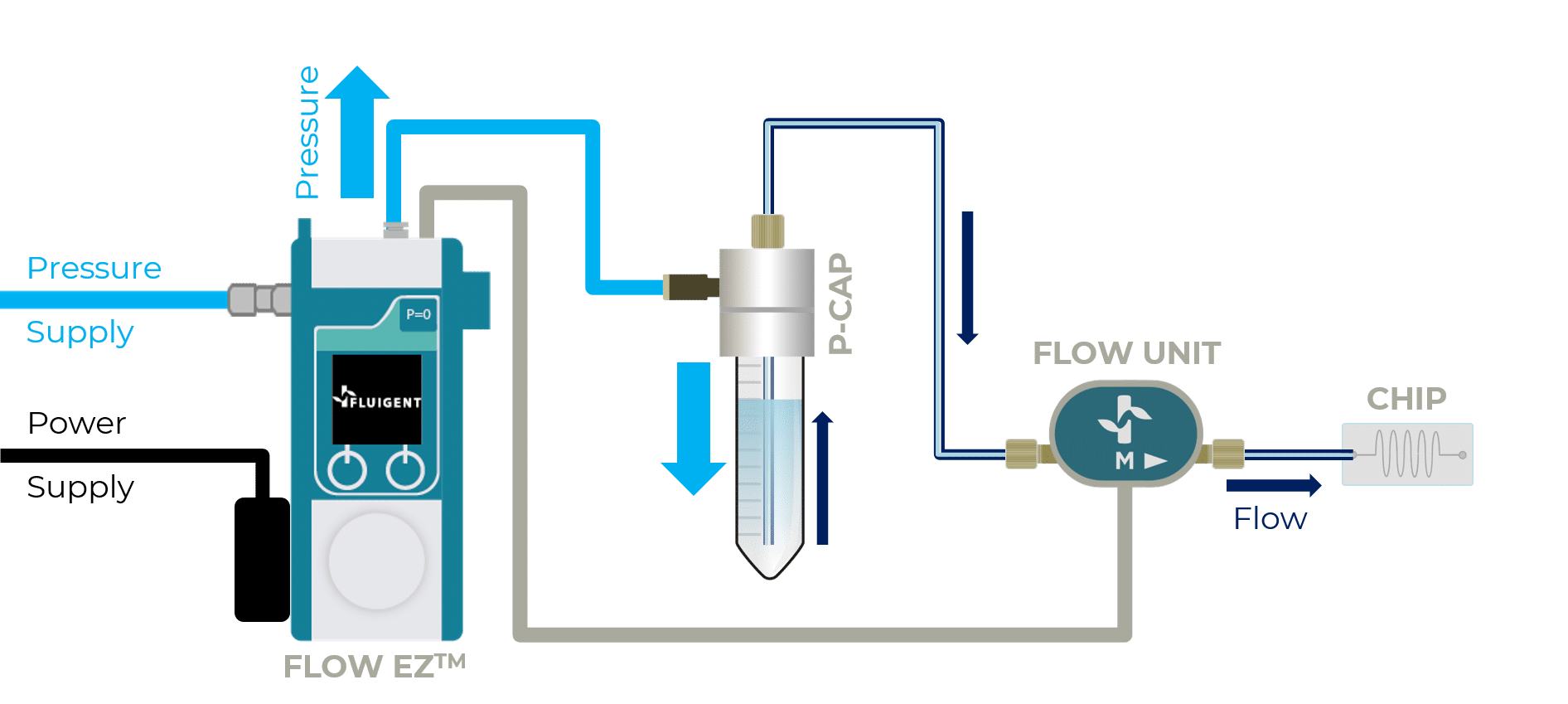
Getting Started
Cell culture
Expertise & resources
-
Microfluidic Application Notes Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions Read more
-
Fluigent Products Datasheets BE-Transflow datasheet Download
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
-
Expert Reviews: Basics of Microfluidics Application of microfluidic chip technology Read more
Related products
For more information on the application access the article:
Direct transfection of clonal organoids in Matrigel microbeads: a promising approach toward organoid-based genetic screens, Nucleic Acid Research, 1–13,
Introduction to Prostate Organoid culture
Advances in three-dimensional (3D) tissue organization models
The past decade has brought significant advances in prostate cancer (PCa) research. With the increased understanding of the origin and the molecular landscape of PCa, there has been an encouraging trend of precision medicine-based approaches to treat advanced PCa.
As we already know, tissues and organs are multicellular structures that self-organize in three dimensions (3D). Cells within a tissue, such as glandular tissue, interact with neighboring cells and the extracellular matrix (ECM) through biochemical and mechanical signals that maintain the specificity and homeostasis of biological tissues.
While traditional 2D cultures on rigid surfaces fail to reproduce cellular behavior in-vivo, 3D matrices are becoming increasingly popular supports for cell culture because they mimic the complex environment that supports the physiological functions of cells to better predict in-vivo responses, thus limiting the need for animal models.
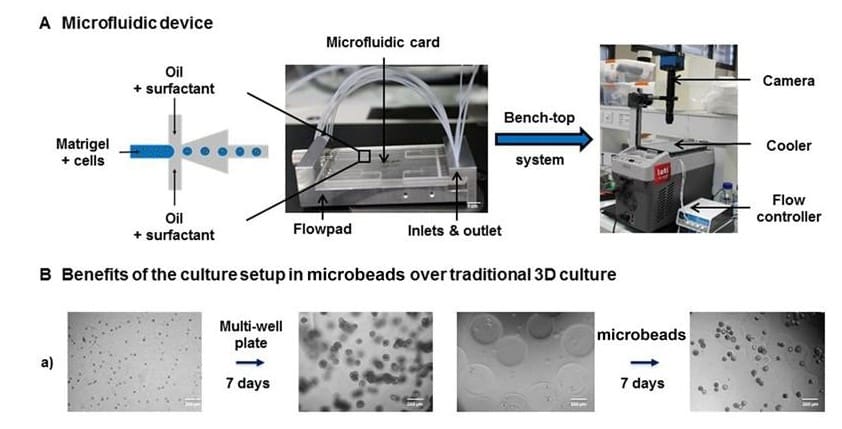
In collaboration with Leti, a technology research institute at CEA Tech.
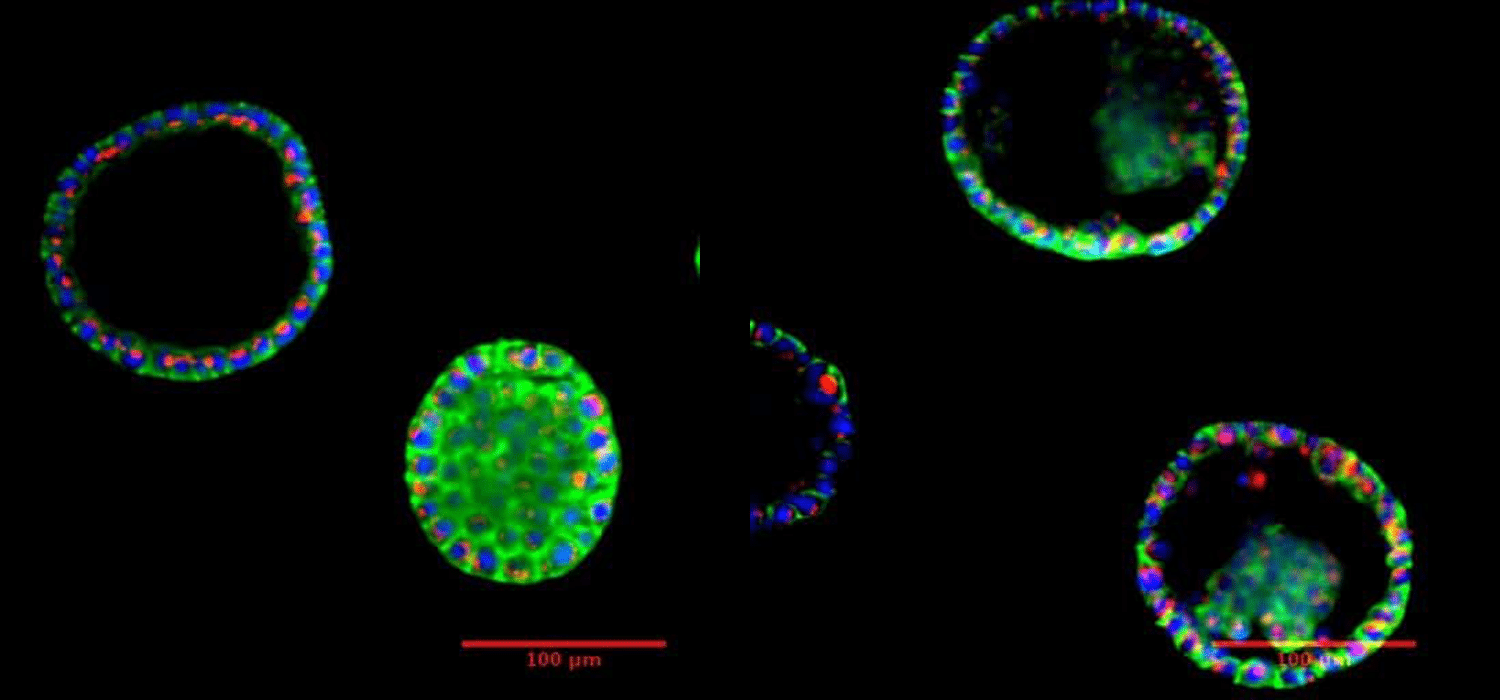
B. Prostate organoid after 7 days incubation. Visualization under fluorescent microscope.
Courtesy of Biomicrotechnology and Functional Genomics (BIOMICS), CEA, Grenoble, FRANCE.
In collaboration with Leti, a technology research institute at CEA Tech.
Organoids as Models for Prostate Cancer Research
Organoids are relevant models to mimic the complex in-vivo environment that supports cell physiological and pathological behaviors. For instance, 3D epithelial organoids recapitulate numerous features of glandular tissues including the development of fully differentiated acini that maintain apicobasal polarity with the hollow lumen. Therefore, researchers have been focusing on the production of prostate organoid culture to better understand the complexity of PCa initiation and progression.
Effective genetic engineering in prostate organoid culture would provide new insights into organogenesis and carcinogenesis, helping us to decipher the key genetic networks underlying epithelial differentiation and polarity, and allowing us to better understand how they may be altered in pathological states such as cancer.
Challenges in 3D Transfection of Organoids
However, direct 3D transfection on already-formed organoids remains challenging. One limitation is that organoids are embedded in the extracellular matrix and grow into compact structures that hinder transfection using traditional techniques. To address this issue, Laperrousaz, B. et al. (2018) have developed an innovative approach for transgene expression in 3D prostate organoid culture by combining single-cell encapsulation in Matrigel microbeads using a Fluigent microfluidic device and electroporation.
Laperrousaz, B. et al. (2018) demonstrated that direct electroporation of encapsulated prostate organoid cultures reach up to 80% of transfection efficiency when combining Fluigent’s technology for organoid generation and efficient 3D transfection. They were also able to validate the role of p63 and PTEN as key genes in acinar development in breast and prostate tissues confirming that this encapsulation and transfection method opens up new perspectives for flow-based high-throughput genetic screening and functional genomic applications.
Benefits of Organoid Culture
Single-cell embedded in microbeads: Each single microbead is considered as a single ‘bioreactor’ for 3D cell culture.
Clonal generation of organoids: Each single encapsulated cell gives rise to an organoid derived from clonal origin.
Monodispersity: The High Throughput (HT) formation of beads (2000 microbeads/min) of controlled size, shape, composition and cell distribution allows for the generation of homogeneous and ‘standardized’ organoids.
Reduced volume of matrix: 2 to 3 times less ECM (ExtraCellular Matrix) than traditional cultures. For example, 350 µl ECM (one well of a LabTek 4-chambers slides) produces 42.800 microbeads with a diameter of 250 µm). This is a great advantage of prostate organoid culture as one of the limitations of these tissues is the simulation of a matrix-embedded environment.
Easy handling: Recovery of organoids in culture media for further analysis.Storage: Microbeads with embedded organoids can be cryopreserved for long periods without altering the architecture and function of organoids.
Storage: Microbeads with embedded organoids can be cryopreserved for long periods without altering the architecture and function of organoids.

| 2D CELL CULTURE | STANDARD 3D CELL CULTURE | 3D CELL CULTURE IN MICROBEADS | |
|---|---|---|---|
| Biological relevance | Low | High | High |
| Control over 3D culture | / | Low | High |
| Easy handling | Yes | No | Yes |
| Clonality | No | No | Yes |
| Transfection efficiency | High | Low | High |
| Long term storage | Yes | No | Yes |
| High throughput | Yes | No | Yes |
| Cost | Low | High | Medium |
Organoid Culture Applications
Functional genomic studies: Controlled organoid generation combined to 3D iRNA-based electroporation in beads opens new perspectives for flow-based HT genetic screening and functional genomic application. As is the case with prostate organoid culture, the transfection efficiency is optimized by modulating microbead size and ECM concentration. The reduced amount of ECM surrounding organoids constitutes a permissive 3D environment that facilitates transfection. PubMed link ».
Tissue development and tumorigenesis: Collecting microbead-containing organoids at different stages allows users to perform a multi-omics analysis of organoid development or carcinogenesis. PubMed link »
Organoids / tumoroid-based drug assays: Flow-based strategies prove to be convenient for future HT Screenings in 3D models and identifying potential RNAi therapeutics. PubMed link »
3D Tool-box: Floating 3D organoids in beads can easily be aspirated, dispensed and sorted by large-particle fluorescence-assisted cell sorting. This flow-based technology opens up broad applications in the field of 3D culture. PubMed link ».


Expertises & Resources
-
Microfluidics White Papers Microfluidic white paper – A review of Organ on Chip Technology Read more
-
Microfluidics White Papers An exploration of Microfluidic technology and fluid handling Read more
-
Microfluidic Application Notes Cancer Cell Analysis Made Easy with Aria: cell Capture and Labeling Read more
-
Microfluidics case studies The Hebrew University: Encapsulation and culture in 3D hydrogels for Single cell sequencing Read more
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
Informative bibliography
[1] Laperrousaz, B., Porte, S., Gerbaud, S., Ville, H., Gidrol, X., Hourtane, V., & Picollet-D’hahan, N. (2018). Direct transfection of clonal organoids in Matrigel microbeads: a promising approach toward organoid-based genetic screens, Nucleic Acid Research, 1–13.
[2] Dolega, M. E., Abeille, F., Picollet-D’hahan, N., & Gidrol, X. (2015). Biomaterials Controlled 3D culture in Matrigel microbeads to analyze clonal acinar development. Biomaterials, 52, 347–357.
[3] Picollet-D’hahan, N., Dolega, M. E., Freida, D., Martin, D. K., & Gidrol, X. (2017). Deciphering Cell Intrinsic Properties: A Key Issue for Robust Organoid Production. Trends in Biotechnology, 35 (11), 1035–1048.
[4] Picollet-D’hahan, N., Dolega, M. E., Liguori, L., Marquette, C., Le Gac, S., Gidrol, X., & Martin, D. K. (2016). A 3D Toolbox to Enhance Physiological Relevance of Human Tissue Models. Trends in Biotechnology, 1–13.
Courtesy of Biomicrotechnology and functional genomics (BIOMICS), CEA, Grenoble, FRANCE.
In collaboration with Leti, a technology research institute at CEA Tech
The Organoids-on-Chip project has received funding from the EU’s H2020 research and innovation program (N°766884) (Read more)
BE-DoubleFlow Features
Easy to use
BEOnChip device is compatible with any type of optical microscopy and its slide format has been chosen for easy handling under a microscope.
Easy to connect
Be-Doubleflow is compatible with all Fluigent pressure-based flow controllers
No unspecific absorption
Unlike in other PDMS devices, the double channel microfluidic chip is made of lipophobic thermoplastic materials and does not present unspecific drug absorption issues. It allows immunohistochemistry with fluorescent detection
Cell recovery
The cell cultures used in the chip can be easily recovered for further experimentation.
Related applications
2D and 3D Culture models used with the BEOnChip device
Crosstalk between neighboring cells underpins many biological processes, including cell signaling, proliferation and differentiation. With this chip, users can explore the crosstalk in the endothelium epithelium barrier under flow in both channels of the device, mimicking a more physiological environment for the study of different organs such as kidney, liver, gut, or heart, among others.
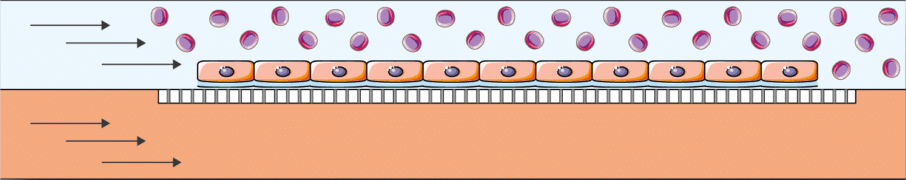
Hypoxic cell culture
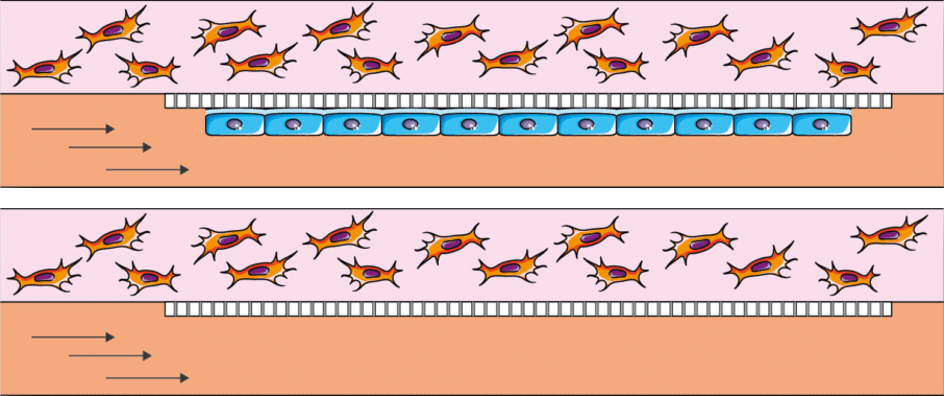
Hypoxia is defined as an insufficient supply of oxygen to body tissues and cells. It plays an important role in various physiological processes and human diseases. As a result, it is an essential subject of experimental study to discover its underlying mechanisms and to create and evaluate treatments for hypoxia-related conditions. As animal models in living organisms cannot fully reproduce certain essential aspects of human physiology, attention has turned to human cell cultures. However, it is important to note that these cultures can introduce biases, particularly when the oxygen concentration around the cells (partial pressure) does not faithfully reproduce the dynamic oxygen conditions found in living organisms. Using the double channel microfluidic chip in combination with Fluigent equipment ensures good mimicking of the in-vivo environment. [1]
In fact, it can combine 3D and 2D cultures in a hypoxia environment where oxygen is supplied to the cells via the medium flowing in one of the channels.
It is also possible to research the effect of circulating particles such as circulating tumor cells, immune system cells, bacteria, fungi, viruses, and many more in single cell culture, or in a complex coculture. For example, the dissemination of circulating tumor cells (CTCs) from primary tumors following metastasis – which then spread to lymph nodes and blood – has been considered an important marker for early cancer diagnosis, demonstrating the importance of studying these CTCs. [2]
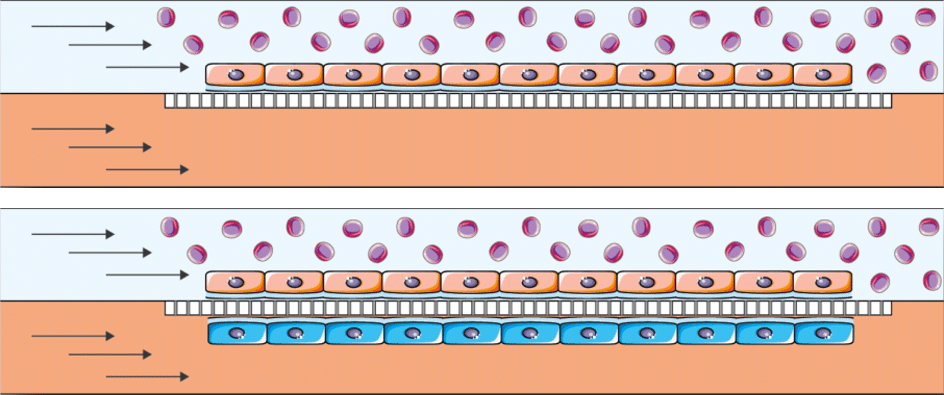
BE-DoubleFlow applications
This chip allows endothelium/epithelium barrier coculture where hypoxic conditions for cell culture are needed or when flux plays a role also in the epithelium culture (Kidney, liver, heart, lung, gut…). In addition, its two perfusable channels offer a perfect environment for studying the effect of circulating particles (bacteria, immune response, circulating tumor cells).
Example applications
Kidney on chip: The coculture of HPTC cells and vascular endothelial cells clearly improves the performance of the HPTCs and increases the maintenance of the renal epithelia in vitro. By using the double channel microfluidic chip, it is possible to coculture both types of cells simultaneously, circulating urine on the renal epithelium culture and blood-like medium in the vascular culture. [3]
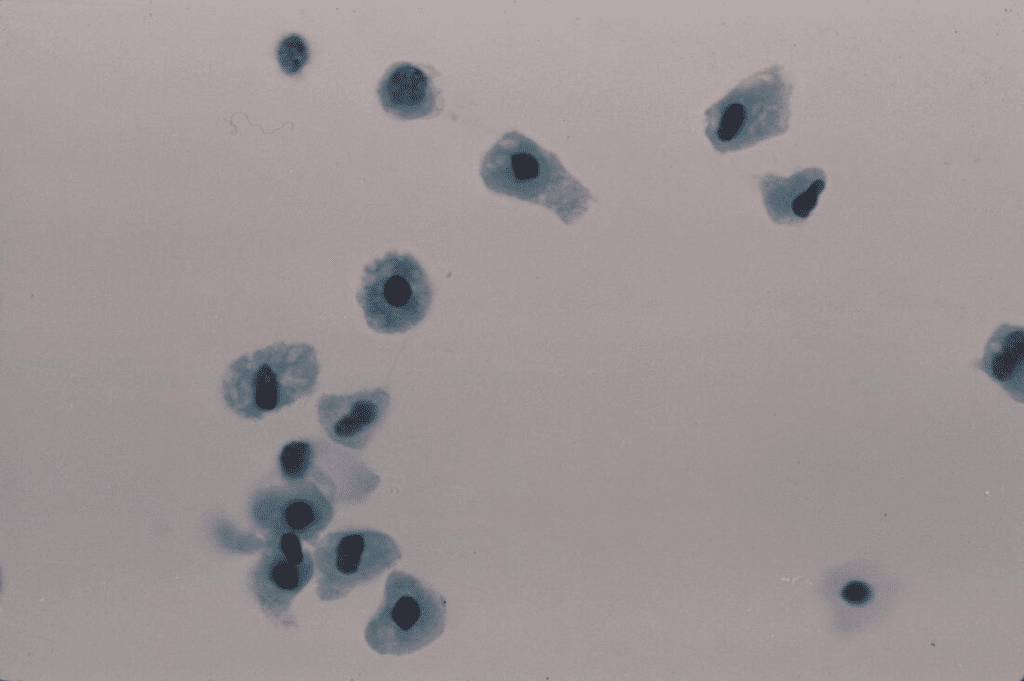
Gut-on-chip: The gut plays a crucial role in fundamental processes such as digestion and absorption, requiring the participation of various cell types. In the context of gut-on-a-chip models, the BEOnChip device is well suited to emulating the low-oxygen conditions typical of anaerobic environments in the microbiome. Moreover, microfluidic methodologies offer the added advantage of controlling the structural arrangement of the cultured model. When it comes to intestinal models, it becomes imperative to assess factors such as the movement of molecules or cells, a task for which conventional 2D well plates are not up to the task. The BE-Doubleflow device provides a solution to this problem, featuring a structure in which two distinct cell culture channels are separated by a porous membrane. This configuration facilitates the cultivation of a variety of cell types, and enables assessments to be made of cell transport and uptake processes. [4]
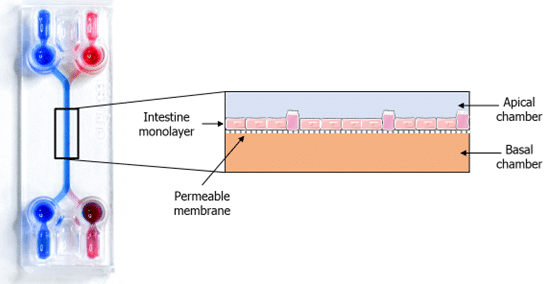
Specifications
PERFORMANCE
| Height | Width | Length | Total volume | |
| Each channel | 375 µm | 1.5 mm | 45 mm | 31.2 µL |
| Inlet/outlet | 7 mm | UNF 1/4″ – 28 | UNF 1/4″ – 28 | 130 µL |
| Medium reservoir | 5 mm | 3.6 mm | 8.8 mm | 185 µL |
| Membrane pore size | 1 µm | 1 µm | 1 µm | 1 µm |

Getting Started
Coating and cell culture
Expertise & resources
-
Microfluidic Application Notes Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications Read more
-
Fluigent Products Datasheets BE-DoubleFlow Standard datasheet Download
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Application of microfluidic chip technology Read more
Reference
[1] Pavlacky J and Polak J (2020) Technical Feasibility and Physiological Relevance of Hypoxic Cell Culture Models. Front. Endocrinol. 11:57. doi: 10.3389/fendo.2020.00057
[2] Das, U.; Banik, S.; Nadumane, S.S.; Chakrabarti, S.; Gopal, D.; Kabekkodu, S.P.; Srisungsitthisunti, P.; Mazumder, N.; Biswas, R. Isolation, Detection and Analysis of Circulating Tumour Cells: A Nanotechnological Bioscope. Pharmaceutics 2023, 15, 280. https://doi.org/10.3390/ pharmaceutics15010280
[3] BEOnChip, BE-DOUBLEFLOW CUSTOM (10 devices per box), BEOnChip Website, Applications section, https://beonchip.com/product/be-doubleflow-custom/
[4] BEOnChip, BE-Doubleflow App. notes: Gut-on-chip 1, https://beonchip.com/be-doubleflow-application-notes-gut-on-chip-1/
Introduction to E. Coli Culture in Droplets
Traditional and new methods for cell culture
Petri plates or culture flasks have traditionally been used in laboratory cell cultivation as part of in-vitro procedures. In culture flasks, cells are grown in a homogenous liquid medium, while Petri dishes allow the growth of colonies on a solid or semi-solid substrate surface. However, these two approaches have limitations, such as the challenge of compartmentalizing clones and single cells.
To overcome these limitations related to culture in Petri dishes or culture flasks, droplet culture of microorganisms, which allows homogeneous growth of cells, is proposed as an alternative method. In this case, E. Coli culture in droplets will be performed. The ability to miniaturize microscale droplets offers great advantages due to their higher surface-to-volume ratio. This feature confers faster mixing and heat transfer, which accelerates reaction times. In addition, droplets are isolated monodisperse chambers that act as reproducible microreactors that can be created with high throughput.
Microfluidics to ensure good monodispersity and stability
However, to be able to use droplets for pharmaceutical and biomedical applications, it is necessary to achieve and guarantee a high degree of monodispersity and stability with such a method. Typically, single emulsions are produced in batches by a two-stage emulsification process (mixing in bulk), resulting in a highly polydisperse population with low encapsulation efficiency.
The use of microfluidics for droplet generation can be a very useful tool to overcome the problems of these conventional methods. In microfluidics, highly reproducible, continuous and reproducible droplet production requires an efficient microchip , a stable fluid handling system (consisting of pulseless pressure-based controllers) and an adapted surfactant.
Why use surfactants in droplet-based microfluidics ?
Surfactants are an essential part of droplet-based microfluidic technology. They are involved in the stabilization of droplet interfaces, in the biocompatibility of the system, and in the process of molecular exchange between droplets. Typically, on encapsulation mechanisms, mineral oils have been the most commonly used with cells. They are therefore limited to the applications such as PCR where the objects of interest (the DNA or RNA fragments) do not exchange between the droplets.
For E. Coli culture in droplets, perfluorinated oils have shown several advantages compared to other carrier fluids such as mineral oils. Their low viscosity allows easy handling in microfluidic systems without need of high-pressure pumps. Though perfluorinated oils have advantages for applications like cell culture and dPCR, it is still difficult to develop a suitable and effective surfactant for droplet stabilization in these oils. To date, most commercially available surfactants present limitations.
The objective of this study is to highlight the biocompatibility of the new surfactant dSURF by performing E. Coli culture in droplets using Fluigent’s pumping technology.
How to perform an E. coli culture
Reagents:
Continuous phase Reagents:
Novec HFE-7500 (Sigma Aldrich) containing 0.5% or 3% dSURF
Dispersed phase Reagents:
500µl suspension of E. coli ECJW922 in TB (Terrific broth) medium with OD600 (optical density) of 0.005 (5 x 106 CFU/ml) or 0.01 (10 x 107 CFU/ml), which means cell concentration in the first suspension was 5 x 106 CFU/ml and in second suspension 10 x 107 CFU/ml. CFU – colony forming unit.
How to generate droplet for encapsulation purpose
E. Coli culture in droplets was performed with customer designed PDMS chip, but we would recommend using the EZ drop chip.
To generate droplets, fluid-handling devices such as pressure controller, syringe pump, or peristaltic pump can be used with a flow-focusing PDMS chip.
Pressure controllers such as Fluigent LineUPTM series or MFCSTM-EZ are best suited to optimize droplet generation performance, improving Flow rate stability and droplet monodispersity. In this case, E Coli suspension and dOIL with dSURF are loaded into vials.
A pressure is applied to the reservoir to ensure a continuous and pulseless injection of both phases into the chip. The determined flow rate is monitored and controled by using OxyGEN software or local control on the Flow EZTM to achieve the desired droplet size and frequency.
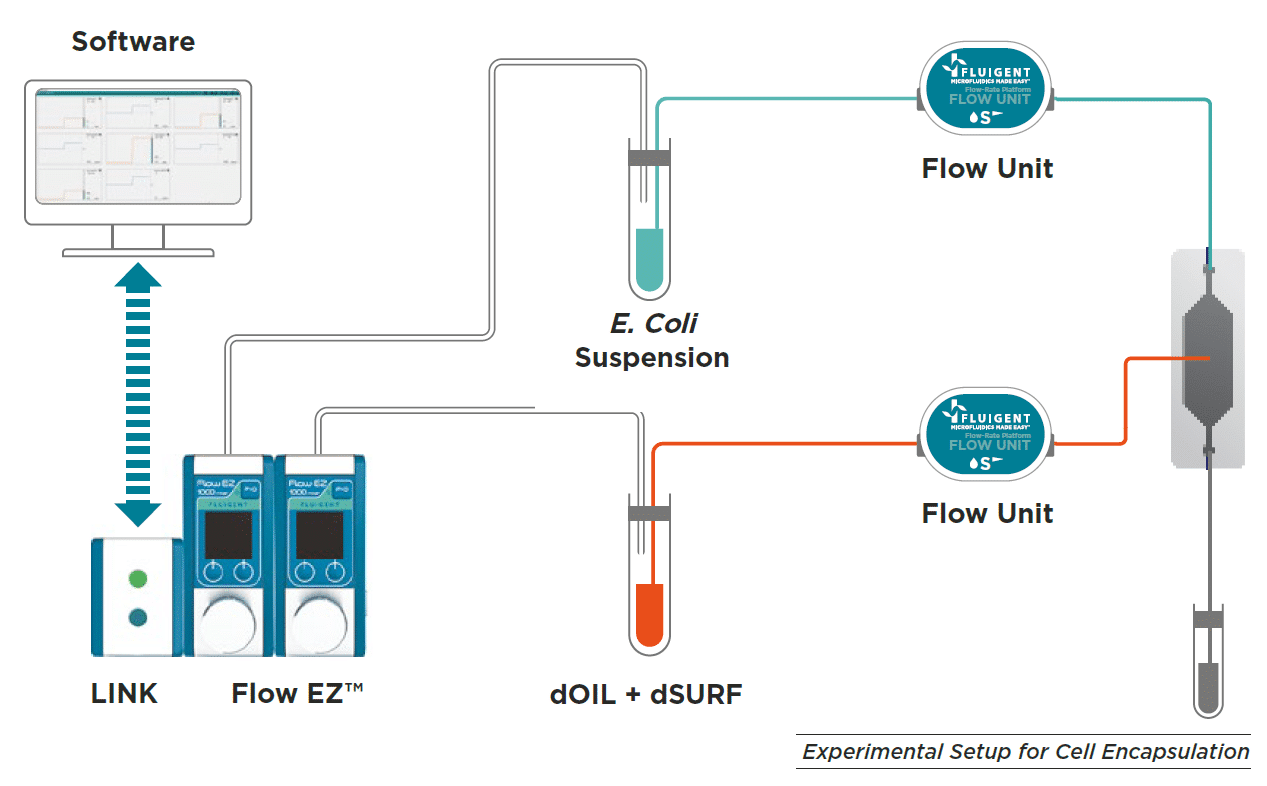
Material for Escherichia coli culture in droplets
Observation of the droplets produced
Test with E. coli cells at OD= 0.005 and 0.5% surfactant concentration, incubation at 37 °C:
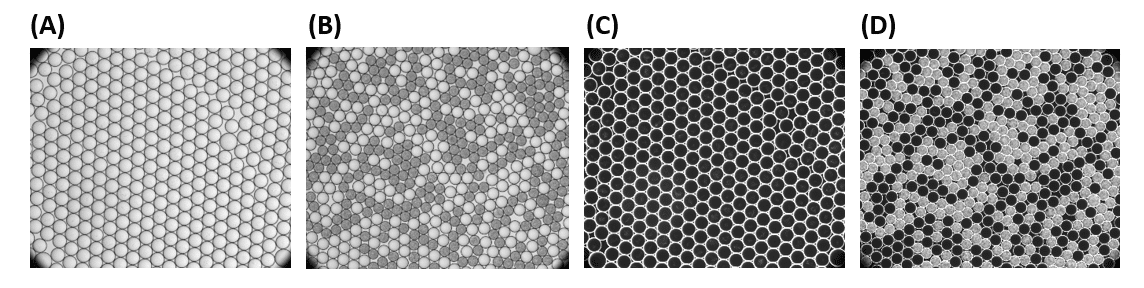

C) Images of droplets under dark-field after 4 h incubation (D) Images of droplets under dark-field after 20 h incubatio
Stable, monodispersed droplet emulsions were observed on both images. High numbers of E. coli cells were observed in almost half of the droplets, particularly in 46±0.8 % of droplets, which is in a good agreement with the theoretical number of 54% calculated from the Poisson distribution of cells over 33.6 µm droplets (160 pL) at a given concentration of 5 x 106 CFU/ml.
The choice to use a low E. coli concentration and fractional droplet occupation with cells, as well as small variations in the experimental process, such as general pipetting imprecision, may help to explain the disparity between theoretical and experimental numbers.
Conclusion
The growth of the E. Coli culture in droplets after 20 h of incubation in all above-described cases has highlighted that dSURF is biocompatible at concentrations ranging from 0.5% to 3%. For routine applications and cost reduction, 0.5% concentration of dSURF can be used. It is also suitable for experiments starting with single and multiple microbial cells per droplet.
The emulsion stability after 20 h shows the good droplet stabilization which will be of use for longer duration experiments.This study has been made in collaboration with Dr Oksana Shvydkiv and her lab from Leibniz Institute for Natural Product Research and Infection Biology.
Expertises & Resources
-
Microfluidics Article Reviews Pressure-driven flow controllers vs. Syringe pumps: A flow precision evaluation for optical blood imaging. Read more
-
Microfluidics White Papers Double emulsion for the generation of microcapsules – a Review Read more
-
Microfluidic Application Notes What is the best method for Microencapsulation of Bacteria and Yeast in Small Double Emulsions? Read more
-
Microfluidics White Papers Droplet-based Microfluidics Read more
-
Expert Reviews: Basics of Microfluidics Microfluidic Droplet Production Method Read more
Features of the microfluidic cell culture chip
Easy to use
The Be-Flow perfused cell culture chip is compatible with all kinds of optical microscopy and has the dimensions of a microscope slide, making it easy to handle under a microscope.
Easy to connect
Be-Flow is compatible with all Fluigent pressure-based flow controllers and can easily be coupled to flow sensors and valves.
No unspecific absorption
In contrast to PDMS devices, there is no non-specific absorption of chemical agents with Be-Flow, allowing for precise control of chemicals inside the chip. In fact, the chip is made of lipophobic thermoplastic materials.
Cell recovery
The cell cultures used in the Be-Flow can be easily recovered for further experimentation.
Related applications
What cell culture models are possible?
Perform 2D culture experiments under flow, controlling mechanical shear stress on the cells in two independent channels.
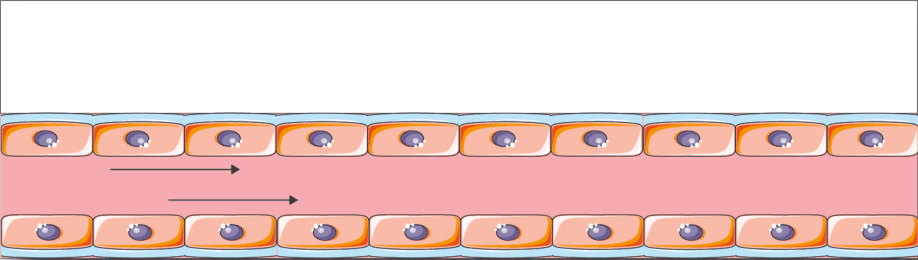

The Be-Flow perfused cell culture chip allows 2D culture on all walls of the device: top, bottom, etc. (see technical note). Co-culture with monolayers of two different cell types is also possible.
Investigate the effect of circulating particles such as circulating tumor cells, immune system cells, bacteria, fungi, viruses, and many more in a 2D culture.
What applications are possible with this microfluidic cell culture device?
BE-Flow is the ideal device for research on the effect of flow and mechanical stress on a cell culture. Some of the most common applications include mechanical shear stress studies, interstitial flow on 3D cultures, rolling and adhesion, or circulating particle experiments.
Vascular research: Because the BE-flow microfluidic cell culture chip was designed for vascular research, some of the most common applications of the BE-Flow device are related to this field of knowledge. Endothelial vascular cells like HUVEC or HAEC are influenced by various forces in the human body due to blood flow. Using the BE-Flow, it is possible to precisely control shear stress and transmural pressure at the same time. These forces play an important role in the gene expression of HUVEC cells and their proper development. In addition, these forces can be modified to a disease level to study phenomena such as thrombosis, atherosclerosis or rolling adhesion of circulating particles in the endothelial tissues.
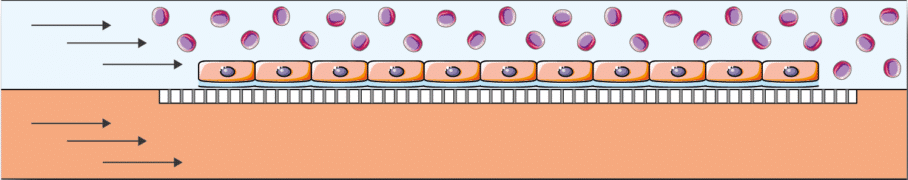
Specifications
Chip dimensions
| Height | Width | Length | Total volume | |
| Each channel | 375 µm | 1.5 mm | 45 mm | 31.2 µL |
| Inlet/outlet | 7 mm | UNF 1/4″ – 28 | UNF 1/4″ – 28 | 130 µL |
| Medium reservoir | 5 mm | 3.6 mm | 8.8 mm | 185 µL |
Getting Started
Coating and cell culture
Expertise & resources
-
Microfluidic Application Notes Long-term fluid recirculation system for Organ-on-a-Chip applications Read more
-
Microfluidic Application Notes Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications Read more
-
Fluigent Products Datasheets BE-flow datasheet Download
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
-
Expert Reviews: Basics of Microfluidics Application of microfluidic chip technology Read more
Related products
What is a flow gradient microfluidic chip ?
A flow gradient chip is a device that combines microfluidics and electrochemical techniques to create controlled and precise biochemical environments for various applications, such as chemical analysis, cell studies, drug testing, and more.
It utilizes electrochemical reactions to create concentration gradients of specific chemical within the microfluidic channels. A concentration gradient is a gradual change in the concentration of a substance across a distance. This gradient is created by introducing different concentrations of the chemicals at different locations on the chip and allowing them to diffuse and mix within the microfluidic channels.
3D cell culture models with gradients
Apply an electrochemical gradient to your 3D cell culture. First mix your cells in a liquid hydrogel and seed them into the central chamber. After hydrogel polymerization has been completed, perfuse culture media with different concentrations of a chemical compound through the lateral channels and monitor the effect in real time.
Be-gradient Features
Easy to Use
The chip is compatible with any type of optical microscopy and its slide format has been chosen for easy handling under a microscope.
Easy to Connect
The BEOnChip device is compatible with all Fluigent pressure-based flow controllers
No Unspecific Absorption
Unlike in other PDMS devices, Be -Gradient is made of lipophobic thermoplastic materials and does not present unspecific drug absorption issues. It allows immunohistochemistry with fluorescent detection
Cell Recovery
The cell cultures used in the chip can be easily recovered for further experimentation.
Applications with a flow gradient microfluidic chip
The flow gradient chip is a microfluidic chip designed with a very specific function in mind, studying a 3D cell culture under an electrochemical gradient. This device allows experiments that can never be done in a petri dish such as the application of nutrient, oxygen or drug gradient, the study of cell migration under these conditions, angiogenesis studies and much more.
Chemotactic migration studies
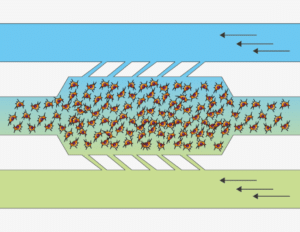
The video shows a Multicellular spheroid embedded in collagen and introduced to the microchamber. A gradient of fetal bovine serum (FBS) was established across the central chamber by addition of growth media containing serum into one of the lateral channels.
We observe that spheroids of oral squamous carcinoma cells OSC–19 invade collectively in the direction of the gradient of FBS. This invasion is more directional and aggressive than that observed for individual cells in the same experimental setup. In contrast to spheroids of OSC–19, U87-MG multicellular spheroids migrate as individual cells.
A study of the exposure of spheroids to the chemoattractant shows that the rate of diffusion into the spheroid is slow and thus, the chemoattractant wave engulfs the spheroid before diffusing through it.
The control of the chemotactic gradient across the microchamber, coupled to the ability to observe and closely monitor the system over time, makes BE-Gradient a powerful technique for the study of the chemotactic process. In fact, 3D matrices can mimic in vivo conditions and thus serve as precious tools for cell migration and chemotaxis studies.
In recent years, collective invasion has been proposed as the dominant migration mode during epithelial tumor development. Using the flow gradient chip enables 3D cell culture process to be observed and analyzed in vitro.
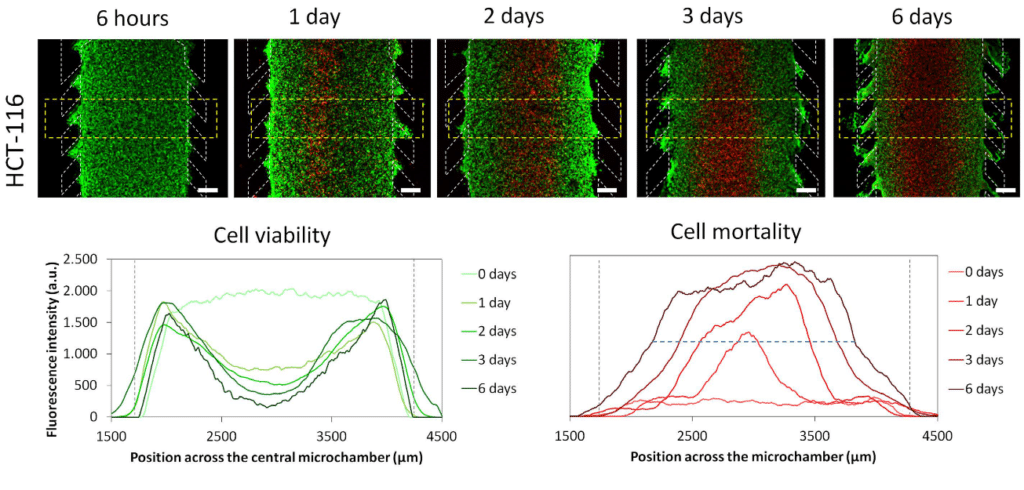
Necrotic core generation within the microdevice
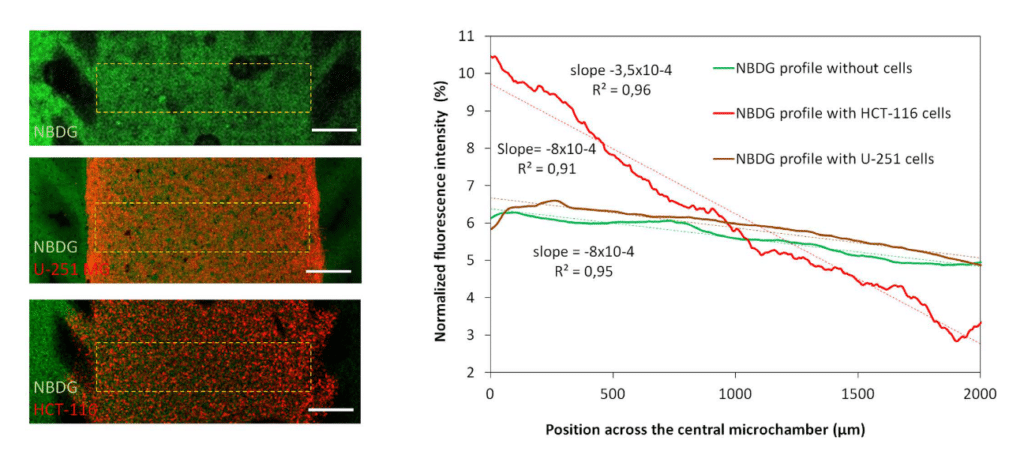
HCT-116 cells were embedded in collagen hydrogel in the central microchamber of the BEOnChip device. 40 million HCT-116 cells/ml were confined in the central microchamber and cell viability was evaluated at the indicated times using calcein (CAM) to stain viable cells green and propidium iodide (PI) to stain dead cells red.
The graphs show CAM or PI fluorescence intensity profile along the delimited region in the images. Position of the pillars is delimited by a grey dashed line. The width of the necrotic core after 6 days was measured as the distance between those positions in the microchamber that reached 50% of the maximum PI fluorescence intensity (blue dashed horizontal line). Necrotic core width was 1643 ± 9 μM, p-value < 0.05. Scale bar is 400 μm.
Glucose gradient
Green fluorescent glucose analogue (NBDG, 200 μ M) was perfused through the left lateral channel of the flow gradient microfluidic chip and the diffusion profile was studied in the absence or presence of cells.
The graph shows the NBDG diffusion profile across the central microchamber after 90 min, demonstrating that NBDG was able to penetrate through the collagen hydrogel. The diffusion profile slope was calculated in the absence of cells or in the presence of HCT-116 or U-251 MG cells. Scale bar is 400 μm.
Flow gradient Chip Specifications
Dimensions
| Height | Width | Length | Total volume | |
| Central channel | 300 µm | 1 mm | 39 mm | 12.6 µL |
| Lateral channel | 300 µm | 1 mm | 50 mm | 14.5 µL |
| Chamber | 300 µm | 2 mm | 4.6 mm | 3 µL |
| Inlet/outler | 8 mm | – | 2.3 mm | 18.4 µL |
| Reservoir | 6 mm | 3.6 m | 7 mm | 151.2 µL |

3D cell culture with gradients
Getting Started
Expertise & resources
-
Microfluidic Application Notes Development of a human gut-on-chip to assess the effect of shear stress on intestinal functions Read more
-
Expert Reviews: Basics of Microfluidics How to choose a microfluidic chip Read more
-
Fluigent Products Datasheets BE-Gradient datasheet Download
-
Expert Reviews: Basics of Microfluidics Passive and active mechanical stimulation in microfluidic systems Read more
-
Expert Reviews: Basics of Microfluidics Mimicking in-vivo environments: biochemical and biomechanical stimulation Read more
Related products

Microfluidic flow controller
Microfluidic flow controller
See the offer
Bidirectional Microfluidic Flow Sensor
Bidirectional Microfluidic Flow Sensor
See the offer
Automated Perfusion System for Spatial Omics
Aria
See the offer
Microfluidic cell culture chip
BE-Flow
See the offer
Double channel Microfluidic chip for hypoxic cell culture
Microfluidic chip for 2D and 3D cultures
See the offer
Air Liquid Interface Cell culture versatile chip
The most versatile cell culture chip
See the offer
Omi, automated organ-on-chip platform
Innovative Organ on A Chip Platform
See the offer
High Throughput Cell Perfusion Pack
High Throughput Cell Perfusion Pack
See the offer
Introduction
Why use a surfactant?
Surfactants are used for various reasons in many industries and applications. Firstly, they lower the surface tension of liquids, allowing better wetting, spreading, and penetration of substances. They can act as dispersants, helping to evenly distribute particles or droplets in solutions.
Additionally, surfactants can modify the interfacial properties between immiscible substances, such as oil and water, enabling processes like emulsification and enhancing oil recovery. Overall, surfactants play a crucial role in improving the efficiency and effectiveness of various industrial and consumer products such as microbiome in droplet experiments.
What is a microbial surfactant?
Microbial surfactants, also known as biosurfactants, are compounds produced by microorganisms that lower surface tension between substances. They possess hydrophilic and hydrophobic regions, enabling interaction with water and non-water substances. These surfactants have diverse compositions, including glycolipids, lipopeptides, phospholipids, and polymeric surfactants.
They offer advantages such as surface tension reduction, foaming ability, environmental compatibility, enhanced oil recovery, antimicrobial properties, and bioactivity. Their applications span agriculture, food processing, pharmaceuticals, cosmetics, bioremediation, and more. Microbial surfactants are an active area of research in biotechnology due to their unique properties and potential uses.
What are the advantages of biosurfactants over chemical surfactants?
Biosurfactants, or microbial surfactants, offer several advantages over chemical surfactants. They are environmentally friendly and derived from renewable resources, ensuring sustainability. Biosurfactants are non-toxic, posing minimal risks to humans and the environment.
They exhibit higher biodegradability, easily breaking down into simpler compounds. Their versatility allows customization for specific applications. Biosurfactants maintain functionality under extreme conditions, making them suitable for various industries. They synergize with microorganisms, enhancing their survival and activity.
Additionally, biosurfactants possess unique properties such as antimicrobial and anti-adhesive effects. While chemical surfactants still have their uses, biosurfactants provide a sustainable and eco-friendly alternative with diverse benefits, that makes it the perfect tool to perform microbiome culture in droplets.
How to Make Droplet-based Microbiome Culture
Reagents
Novec HFE7500 (3M) contains 5%, 2% or 0.5% (w/w) concentration of Competitor 1 surfactant, Competitor 2 Surfactant, or dSurf (Fluigent) .
Microbial sample
A microbial sample collected from the skin of healthy volunteers was used for microbiome culture in droplets generation. A general microbial skin sample, as opposed to a single strain like E. coli, was used here because a bacterial community consisting of many species better represents the experimental condition for microbiological research.
The microbial skin sample was prepared according to Biomillenia’s proprietary standard sample preparation workflow and was suspended in a standard medium for skin microbes containing 0.5% (v/v) TWEEN80. TWEEN80 is a lipophilic molecule in the aqueous phase that interferes with the droplet stabilizing characteristics of the surfactants.
Droplet generation
The droplets were prepared on Biomillenia’s proprietary microfluidic platform with PDMS chips. The two liquid phases were controlled during droplet generation by the use of high precision pressure pumps (Fluigent, Flow EZ).
Droplets were generated at frequencies of 7-10 kHz. The droplet volume was set to 20 pL. Droplets were collected and incubated in custom made vessels at 37 °C, allowing the droplets to be stored without exposing them to a direct gas interface.
Observation of the microbiome culture in droplets
From the droplet collection vessel, a small number of droplets were sampled to check for droplet stability and bacterial occupation at 0, 1, 3 and 7 days of incubation. For imaging by microscope, droplets were spread onto a monolayer surrounded by an oil-surfactant combination.
What material to perform droplet microfluidics for microbiome culture?
Partial results of droplet-based microbiome culture
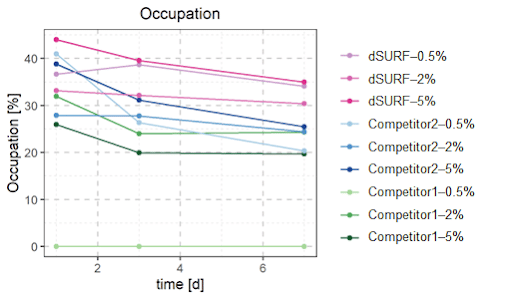
The collected droplets were spread in Biomillenia’s proprietary observation chip for best imaging of microbial occupation in droplets after 7 days of incubation at 37 °C. As shown in the figure below, a monolayer of droplets surrounded by the respective oil-surfactant combination was imaged microscopically.
Surfactant concentrations of 2% and higher result in stable populations over 7 days at 37°C for dSURF and Competitor 2. While for Competitor 1,a concentration of 5% is required to reach similar results.
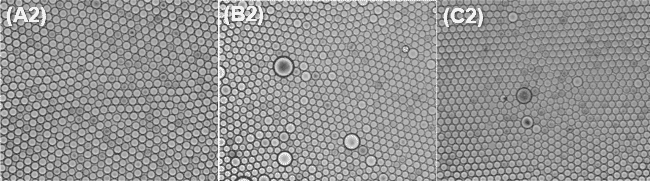

Bacterial occupancy was observed in the microbiome culture in droplets after 1, 3 and 7 days for each condition. A droplet is considered ‘occupied’ when it contains at least 3 bacteria. Since all the bacterial encapsulations were done in the same condition (same bacterial culture at the same concentration, same flow-rates, etc), one would expect the bacterial occupancy to be similar for each condition.
However, the choice of surfactant has a strong impact on the occupation rate, meaning a higher and more stable occupation rate is found with dSURF surfactants.
Conclusion
The stability of microfluidic emulsions strongly depend on the content of droplets and its interplay with the surfactant used. Hydrophobic compounds in the aqueous phase, along with microbial growth of various species, present far from ideal conditions for microfluidic droplets, but exemplify the experimental challenges encountered when microfluidic techniques are applied in microbiology. Hence, surfactants are needed to accommodate those complex sample characteristics.
In conclusion, dSURF surfactants for microbiome culture in droplets perform well compared to widely established microfluidic surfactants and are highly suitable for complex biological applications.
Expertises & Resources
-
Microfluidics case studies Using dSurf for High Throughput Laser-Induced Fluorescence Droplet Micro-Thermometry (LuMIn) Read more
-
Microfluidic Application Notes A quick and efficient double encapsulation method for FACS-based droplet sorting Read more
-
Microfluidics White Papers Droplet-based Microfluidics Read more
-
Microfluidic Application Notes E. Coli Culture in Droplets Using dSURF Fluorosurfactant Read more
-
Microfluidic Application Notes Analysis of a commercial surfactant for digital PCR assay Read more
“Microfluidics is a robust and cost-effective technology for single cell and single clone processing”
“Fluigent made microfluidics parallelization a feasible task”
CloneSeq: A highly sensitive single-cell analysis platform for the comprehensive characterization of cells from 3D culture
Introduction
Single-cell studies have revealed that there is considerable cell-to-cell variation within tumors of different cancer types and during embryonic stem cell (ESC) differentiation 1–4. However, in many cases, single cell experimental data is still difficult to interpret as a high degree of randomness persists.
To overcome this limitation, Bavli et al. developed a complementary single cell sequencing technology: CloneSeq. This method combines clonal expansion inside three-dimensional (3D) hydrogel spheres and droplet-based RNA sequencing (RNA-seq)5. The authors revealed the presence of novel cancer-specific subpopulations, including cancer stem-like cells, which are not identified using standard RNA sequencing assays.
Barcode formation, cell encapsulation in hydrogels, and single cell sequencing (including InDrops and Drop-seq) were all performed using Fluigent Flow-EZ pressure-based flow controllers, controlled with Fluigent OxyGEN software with pressures ranging from 69 mbar to 2 bar.

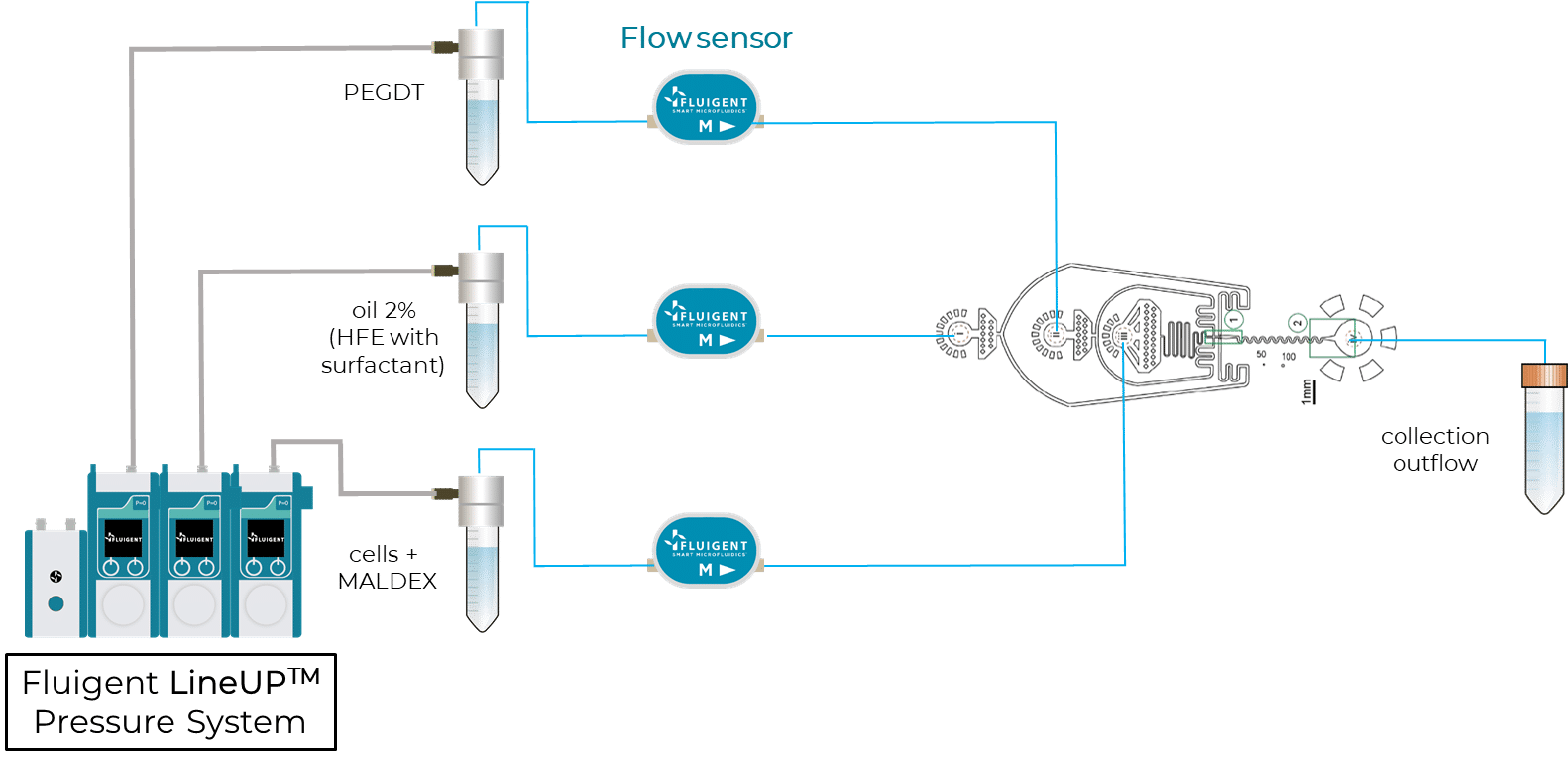
Step 1 – Single-cell encapsulation in hydrogels for 3D cell culture system
The first step of the CloneSeq method consists of capturing single cells inside a biodegradable hydrogel for subsequent clonal expansion. All flow rates are finely controlled using Fluigent Flow EZs combined with our Flow Units and OxyGEN software. Flow rates used range from 8 to 34 µL/min depending on the solution. About 700 hydrogel spheres per second are generated under these parameters5.
The cured gels are collected into a tube and immersed in culture medium, permitting the proliferation of the encapsulated cells and clone formation (figure 2).
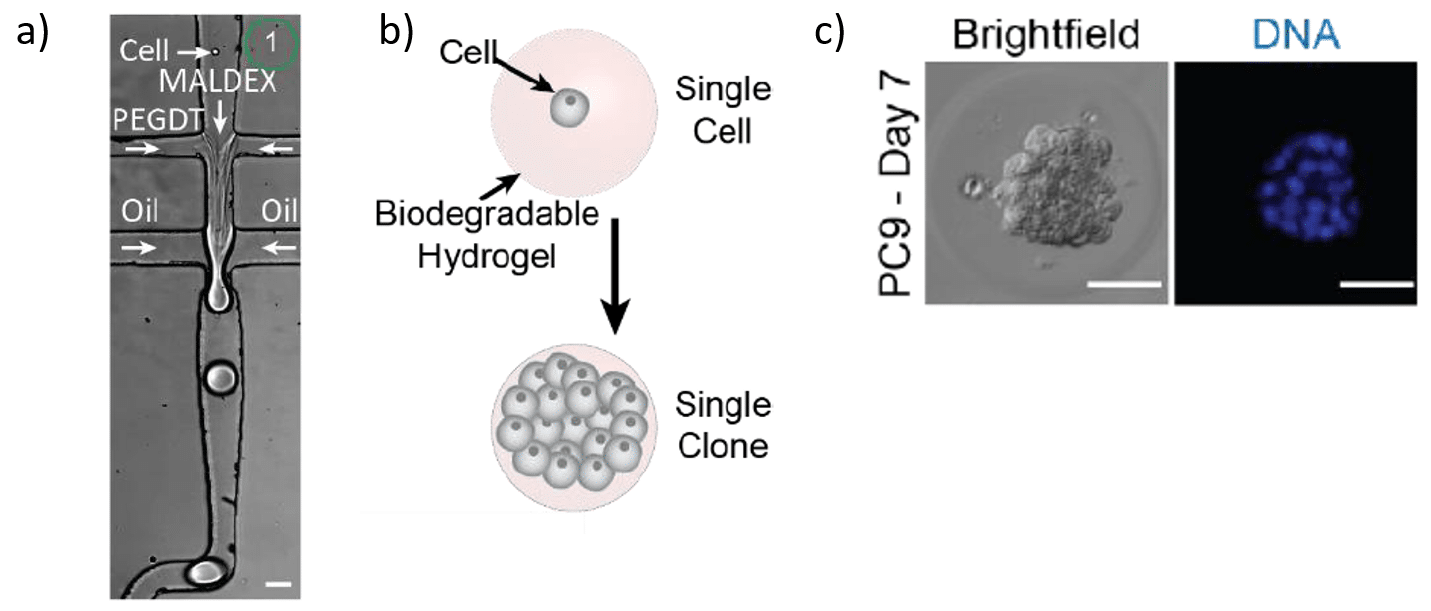
Step2 – CloneSeq: profiling of clones using modified InDrops and Drop-Seq protocols
For the RNA profiling of clones, the authors designed a microfluidic device to capture clones in droplets and barcode their mRNAs using a custom InDrops protocol5. Figure 3 shows a scheme of this InDrop-based microfluidic configuration. Once again, flow rates are finely controlled using Fluigent Flow EZs combined with our Flow Units and OxyGEN.
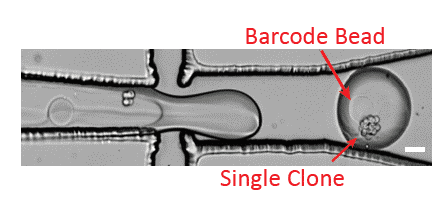
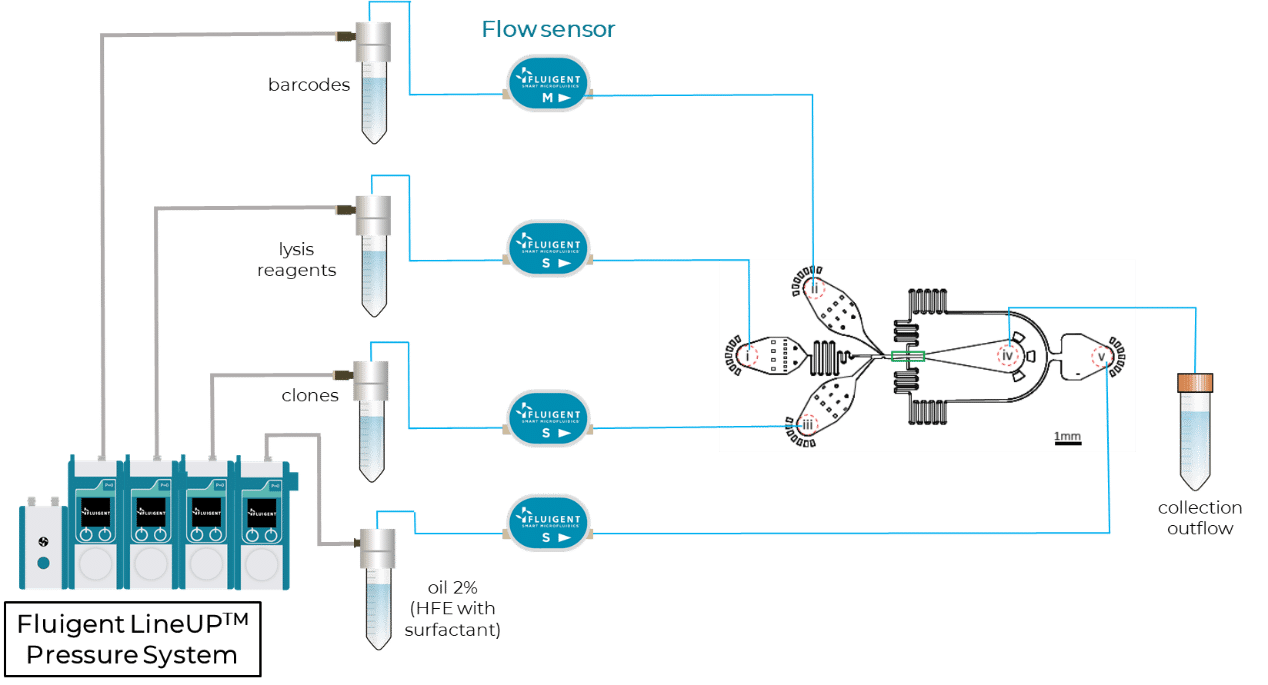

Partial results
To study the effect that clonal expansion within the hydrogel spheres has on the homogeneity of cell states within the clones with droplet-based single cell sequencing, the authors compared the inter-clone correlations of small (n<15 cells) and large (n≥15 cells) clones and of pseudo-clones. They showed for human PC9 cells that cells sharing a clonal origin are more similar to each other compared to random cells, suggesting that clones are homogeneous5.
Cleaning protocol
After the experiments, the authors perform an automated cleaning protocol using the Fluigent M-Switch, a microfluidic valve allowing for the injection of up to 10 fluids in a sequential manner.
Conclusion
With this case study, the authors demonstrated the use of Fluigent pressure-driven flow controllers along with a dedicated software to provide excellent flow control for cutting-edge microfluidic applications including single-cell encapsulation and culture within 3D hydrogels droplets, and for InDrops protocols followed by RNA sequencing.
Related Products
Related Resources
References
- Li, H. et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49, 708–718 (2017).
- Dalerba, P. et al. Single-cell dissection of transcriptional heterogeneity in human colon tumors. Nat. Biotechnol. 29, 1120–1127 (2011).
- Kim, K. T. et al. Single-cell mRNA sequencing identifies subclonal heterogeneity in anti-cancer drug responses of lung adenocarcinoma cells. Genome Biol. 16, 1–15 (2015).
- Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science (80-. ). 352, 189–196 (2016).
- Bavli, D. et al. CloneSeq: A highly sensitive analysis platform for the characterization of 3D-cultured single-cell-derived clones. Dev. Cell (2021).
Oren Lab website & other research articles
https://ramoren3.wixsite.com/oren-ram-lab
https://www.bio.huji.ac.il/en/departments_biological_chemistry_en
Esrrb is a cell cycle dependent priming factor balancing between pluripotency and differentiation
Introduction
Erasmus MC is a leading international academic hospital at the forefront of the medical field. Its staff, volunteers, and students collaborate to provide healthcare for patients with complex disorders, rare conditions, and urgent medical needs. Recognized as a world-class scientific research organization, Erasmus MC strives to improve their understanding of diseases and disorders, working towards prediction, treatment, and prevention. The institution’s guiding principle revolves around the integration of biomedical research, clinical research, and health sciences to form a comprehensive approach to the advancement of healthcare.
Bi/ond, an innovative and international biotechnology company based in the Netherlands, was created with the primary aim of harnessing the potential of microchips to drive biological innovation. Working in close partnership with biologists, the company focuses on the creation of reproducible and accurate biological models that help develop inclusive and precise cures for medical issues. By drawing on its expertise in microelectronics and its in-depth knowledge of biological solutions, Bi/ond succeeds in bridging the gap between the fields of biology and engineering.
These two entities have combined their knowledge and technologies to develop an innovative Cancer-on-Chip (CoC) platform for assessing response to treatments using Fluigent’s high throughput cell perfusion pack.
Learn more about the Erasmus MC Cancer Institute
Learn more about Bi/ond

Testimonial

“We started out as a novice to the field of organ-on-chip cultures. Fluigent was very helpful at this stage to get started and keep the system up and running. They were always there to answer questions. Therefore, the precision pumping system was a thing we did not have to worry about, and we could concentrate on our own expertise, the biological materials in the culture device.”
Dr. D.C. (Dik) van Gent PhD
Universitair Hoofd Docent – Molecular Genetics
Why develop a Cancer-on-Chip (CoC) device to predict drug response?
Overcome the challenges in cancer treatments
Searching for personalized therapy treatment for individual patients is a challenging process. The crux of the problem lies in precisely defining the optimal treatment regimen for each individual. Although many molecular biomarker-based treatment strategies have been employed in cancer therapy, their ability to reliably predict individual responses to chemotherapy remains limited in most cases. This means there is a pressing need for ex vivo bioassays capable of effectively predicting a patient’s response to specific treatments, thus facilitating the selection of the most appropriate and effective therapeutic approach to optimize life expectancy and quality of life.
Historically, cancer cell lines and animal models have played a key role in assessing the efficacy of chemotherapies. However, when it comes to predicting tumor sensitivity in individual patients, these preclinical models fall short. Their main utility lies in studying the general characteristics of specific tumor types or stages of disease, which fails to consider the heterogeneity found in cancer and undermines their predictive power for responses to individualized treatment. These models are also time-consuming to set up and use, which limits their usefulness.
One potential solution lies in the direct assessment of drug responses using patient tumor tissue slice cultures. These ex-vivo cultures maintain the entire tumor microenvironment, including immune cells, and preserve the original tissue’s architecture. However, the development of long-term ex vivo culture systems, particularly those lasting more than 7 days, remains a major hurdle. The difficulty lies in managing mechanical stress on tissue slices, which can lead to the disruption of tissue integrity and non-physiological behavior, as well as in ensuring optimal culture conditions.
Consequently, it is crucial to invest in the development of more physiologically relevant ex vivo tissue slice culture systems, such as a cancer-on-chip platform, which enables the prolonged culture of tumor slices under precisely controlled conditions. Such advances could revolutionize personalized medicine and dramatically improve cancer treatment by enabling more precise and effective therapies, tailored to each patient’s specific needs.
The development of the organ-on-chip platform, an innovative technology
Despite considerable progress in computational and in vitro biology and toxicology over the past two decades, the failure rate of experimental drugs in clinical trials remains high, with over 80% of drugs failing to reach the market. Of these failures, 60% are attributed to lack of efficacy and 30% to toxicity. This situation has given rise to growing concerns about rising costs, wasted time, and ethical problems associated with animal experimentation, which often proves inadequate for predicting human reactions in a clinical context.
In addition, traditional live-cell experiments using cells grown on 2D substrates coated with serum or extracellular matrix molecules present limitations. Although they promote cell proliferation, they often fail to reproduce tissue-specific functions. Human organs, with their diverse functions, rely heavily on complex interactions between specialized cell types at well-defined interfaces, arranged in complex geometries and responding to specific microenvironments. Due to these issues, there is an urgent need for new modeling and testing platforms capable of better predicting human responses.
Organ-on-a-chip (OoC) technology represents a significant advancement in drug discovery and development, offering new tools for disease modeling and characterization, as well as potentially more accurate methods for assessing the toxicity and efficacy of new compounds and therapies. The organ-on-a-chip concept involves reproducing the functions of human physiology or disease at organ level in microfluidic chips, using different cell types.
Microfluidics plays a crucial role in enabling precise control of the cellular microenvironment, presenting cells with mechanical and biochemical signals in a more physiologically relevant context. Working with liquid volumes in the microliter range, these models enable dynamic scaling and interaction between cells. In addition, microfluidic chips can use geometries and structures to mimic physiological length scales with concentration gradients and mechanical forces generated by fluid flow, thus recreating the in vivo microenvironment faced by cells. This biomimetic approach, involved in the cancer-on-chip platform used in this study, overcomes many of the limitations encountered with conventional tissue culture models. [1]
How to combine OOAC and therapy assessments
The CoC platform provides continuous media perfusion, nutrient supply, waste removal, and the ability to collect samples for analysis. The aim of this platform is to develop a reproducible culture system for assessing the sensitivity of (breast and prostate) tumors to chemotherapy, using living material that closely resembles the original tumor and allows long-term culture without significant changes in viability or tissue characteristics. In addition, the system must enable a direct assessment of response to treatment through microscopic imaging and analysis based on fluid sampling.
In this article, the Cancer-on-a-Chip (CoC) microfluidic platform described uses an 6-well plate with silicon-based microfluidic chips. These chips offer greater flexibility than glass-based culture systems, as they allow for the easy integration of sensors for pH detection, metabolite screening, and oxygen sensing. In addition, the silicon-based design enables parallelization, taking advantage of semiconductor technology to improve scalability, reproducibility, and cost-effective large-scale production.
One notable application of this new Cancer-on-a-chip platform is personalized medicine. It facilitates the in vitro culture of tumor tissue slices under precisely controlled conditions, enabling the prediction of in vivo tumor responses to therapy in individual patients. The platform has been successfully used to grow tumor slices, including patient-derived xenografts (PDX), and faithfully mimicked tumor cells.
Use of the high throughput cell perfusion pack in the Coc platform to predict drug responses
Mimicking the in vivo cancer tissue on a microfluidic chip
In this study, the microfluidic chips from Bi/ond are made of polydimethylsiloxane (PDMS) film with embedded microfluidic channels, supported by a silicon (Si) frame. The chips’ top plates serve as an interface connecting the inlet and outlet, enabling media diffusion through the tissue-supporting membrane. The chip features four microfluidic fittings for the external pumping systems. The bottom part of the chip is designed to ensure compatibility with microscopes and oxygenation, achieved through PDMS window openings under the chips.
The microfluidic chips containing tumor tissue slices are housed in a ComPLATETM. The ComPLATETM, designed by Bi/ond, is a smart, compact, and reusable well-plate specifically tailored for cultivating complex tissues. The plate is comprised of a black 6-wells bottom plate, a transparent top plate to cover the chip, and a white fixation ring. This design helps create independent cultivation and the analysis of individual tumor slices.
The top plate of the ComPLATETM provides the option for single or double flow of media. Each well has sufficient space to accommodate the tissue slice’s growth over time. The microchannel and top interface facilitate constant perfusion and nutrient replenishment, enabling maintenance of the tissue slices while removing waste products. Furthermore, oxygenation of the tissue slices is enhanced through a gas exchange via the PDMS layer of the optical window.To monitor fluid flow rates inside each well throughout the culture period, the entire Cancer-on-a-Chip platform is connected to a Fluigent Microfluidic Flow Control System.
A, Top view of the microfluidic chip illustrating its components: the PDMS film in which the microfluidics are embedded, and the silicon frame, which includes the inlet and outlet to the channels in the film. B, Vertical cross-section of the microfluidic chip. C, Representation of the CoC platform. D, ComPLATETM device. E, Cross-section of CoC illustrating the diffusion and perfusion toward the tissue slice.
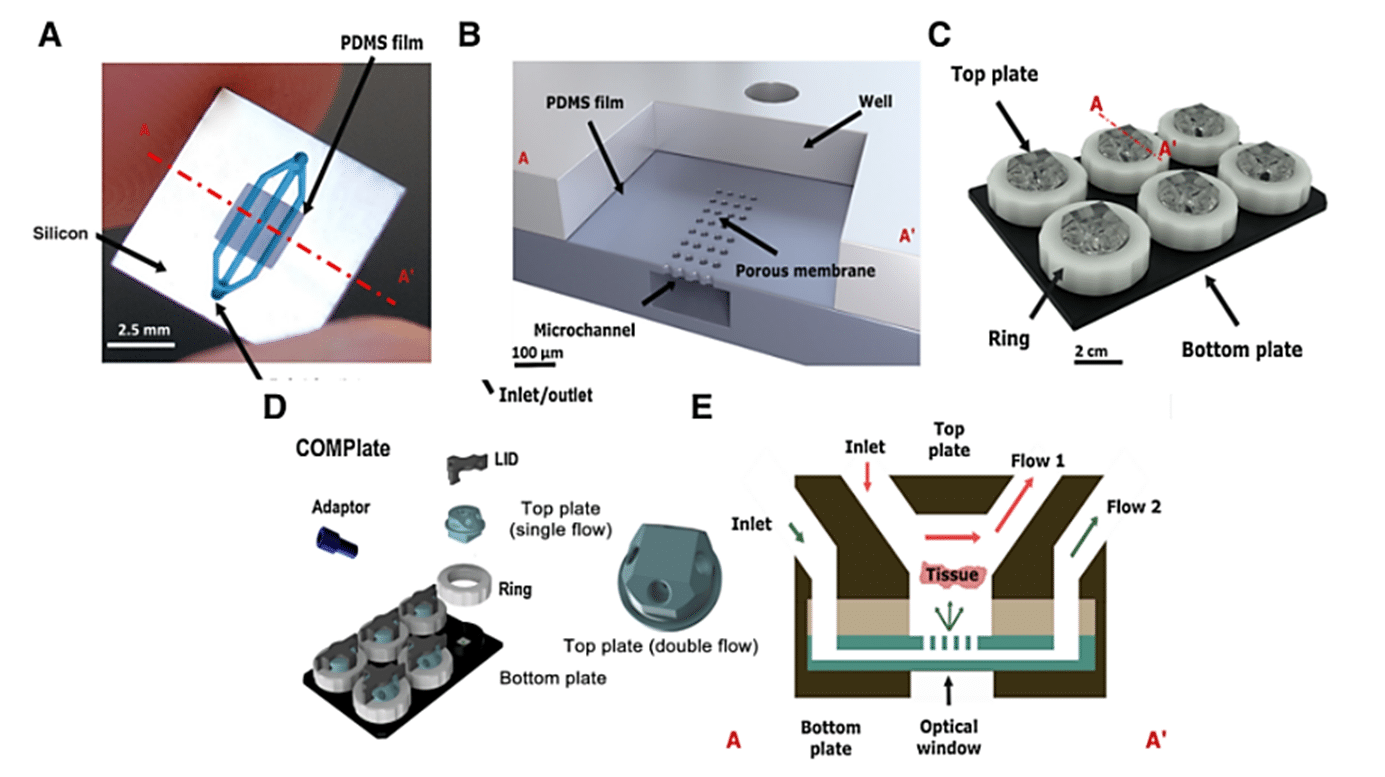
Using Fluigent technology to ensure a high throughput cell perfusion
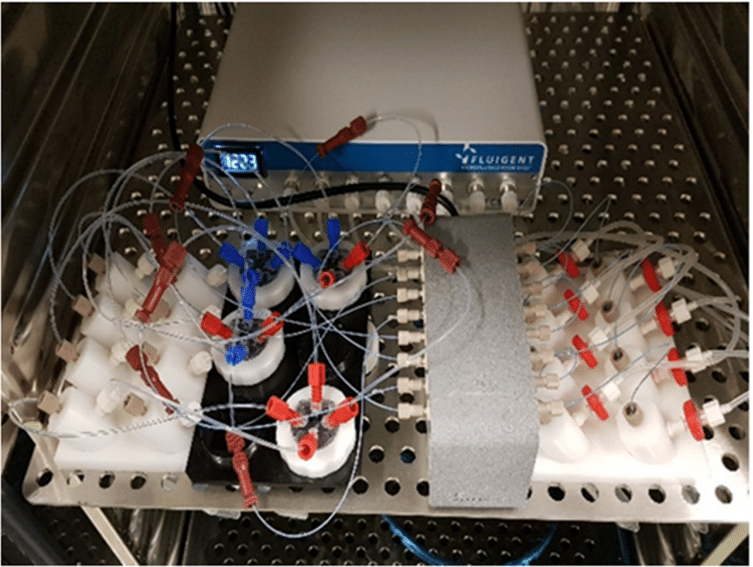
The Fluigent Cell Perfusion Pack is specially designed for high-throughput experiments. It has been carefully optimized for maximum efficiency in multiple-chip perfusions, enabling the simultaneous growth of multiple organ models in one incubator, which is ideal for the cancer-on-a-chip platform. The package includes a compact 8-channel pressure controller, a flow platform, and reservoir support that can be easily integrated into an incubator track. The user-friendly interface makes it easy to set up and operate the system, creating the proper physiological conditions for effortless long-term experiments. The system is highly reproducible and scalable.
To achieve continuous perfusion, an FLPG Plus pumping system was used as the pressure source. The flow rate was then maintained using the MFCSTM-EZ pressure-controlled microfluidic flow control system. Flow sensors (FLOW UNIT-S) were used to monitor flow throughout the culture using Fluigent software. A precise inlet flow rate of 5 µl/minute was used to perfuse PDX tissue slices through the chip’s upper and lower channels. CoC tissue was cultured under optimal conditions in a humidified atmosphere with 5% CO2 at 37°C, and the culture medium was renewed every 3 days for up to 2 weeks.

Comparison with an ex-vivo model
One of the aims of this study is to compare this innovative cancer-on-chip platform with a more traditional ex-vitro model. This traditional model consists of a cell culture in a 3mL customized culture with medium 6-well standard plates on an orbital shaker at 60 rpm. This will demonstrate the added value of using a platform based on Organ-On-a-Chip technology compared to more conventional methods. This will be made possible by carrying out various tests and comparing the data obtained by these two methods. These tests consist of studying the treatment response of tumors, long-term tumor tissue slice culture, and gene expression analysis.
Partial results
Treatment response of tumors
To assess the validity of the platform, a crucial question is whether in vivo treatment responses can be predicted by treatment responses. This validation was carried out using the cisplatin treatment on PDX breast cancer tumors (cisplatin-sensitive and cisplatin-resistant) with three biological replicates each.
To verify that the platform retained the essential features of tumor-associated cell morphology and proliferative capacity, untreated tumor slices at day 0 and day 7 were evaluated.
The effect of cisplatin treatment on cell proliferation and death was assessed in tissue slices grown under normal ex vivo conditions and in the CoC device. Cisplatin-sensitive tumor slices in the platform showed a significant increase in apoptotic cells and a notable decrease in replicating cells upon cisplatin treatment. In contrast, cisplatin-resistant PDX tissue slices showed no significant changes in either signal compared with untreated controls. [2]
The response to cisplatin treatment observed in the Cancer-on-a-Chip platform correlates with known tumor responses in in vivo and ex vivo cultures, suggesting its reliability for drug response analysis. Interestingly, breast PDXs cultured in the CoC platform showed a more robust response to cisplatin treatment compared with the ex vivo culture method, indicating better drug delivery in tumor slices with the platform.
To evaluate the performance of the Cancer-on-chip device in another tumor type, the PC82 androgen-dependent prostate tumor was used under the same conditions. The results led to the same conclusion as the breast tumor slices. [3]
In conclusion, CoC cultures accurately reproduced tumor responses to two different treatments (prostate and breast) in breast and prostate tumor models known to be sensitive in vivo.
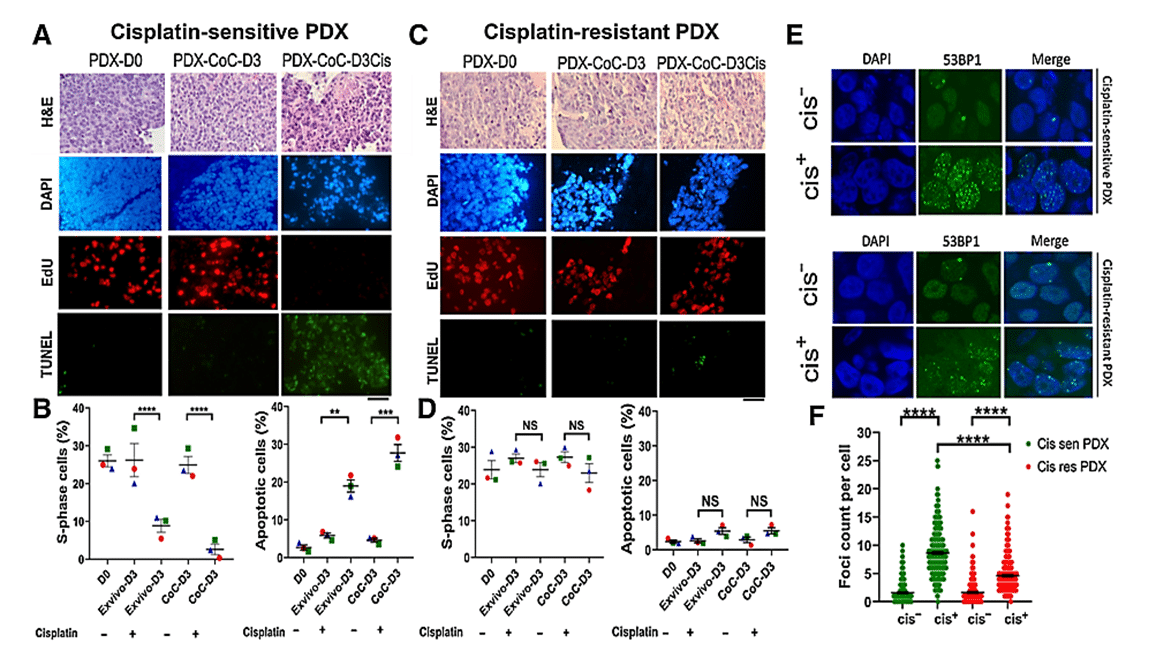
Figure 3: Prediction of therapy response using cisplatin-sensitive and -resistant PDX in ex vivo and CoC platforms. A, Representative EdU (proliferation) and TUNEL (apoptose) staining of cisplatin-sensitive breast PDX. B, Quantification of the fraction of EdU-positive and TUNEL-positive cells showing breast PDXs were sensitive to cisplatin. C, Representative EdU and TUNEL staining of cisplatin resistant breast PDX. D, Quantification of the fraction of EdU-positive and TUNEL-positive cells showing breast PDXs were insensitive to cisplatin therapy, thereby validating the application of CoC for therapy response for patient tumors. E, Analysis of DNA damage response in cisplatin-sensitive and -resistant PDX treated with cisplatin. Cisplatin treatment induced more double-strand breaks in cisplatin-sensitive PDX than in cisplatin-resistant PDX. F, Scatter plot showing 53BP1 foci count per cell in cisplatin-sensitive and -resistant PDX.
Long-term tumor tissue slice culture
Longer culture times are essential for studying therapeutic responses that require longer incubation periods-more than one week-such as the development of therapeutic resistance or clonal outgrowth. The main limitation observed in ex vivo culture is the preservation of optimal proliferative capacity and tissue architecture.
To address this, a study was carried out using breast tumor slices from five independent patient-derived xenografts. These slices were cultured for 14 days in the Cancer-on-a-Chip platform and, in parallel, in the ex vivo 6-well plate. After 7 days of culture, similar rates of cell proliferation in the CoC device compared with day 0 were observed, but slightly slower proliferation in the ex vivo condition. TUNEL staining revealed a slight increase in cell death in the ex vivo system compared with the CoC platform at day 7.
Notably, these differences became more pronounced in extended 14-day cultures. The platform showed better preservation of tumor tissue architecture and cell proliferation than the ex vivo culture system. In contrast, the ex vivo condition showed a significant decrease in proliferation at day 7 compared with PDX tumors at day 0, whereas no significant difference was observed between day 7 and day 0 for the platform, indicating slightly slower cell proliferation in ex vivo culture compared with CoC.
These results underline the superiority of the Cancer-on-chip device for prolonged culture times (beyond 7 days) of tumor tissue slices compared to the ex vivo system (better preservation of cell integrity and cell proliferation).
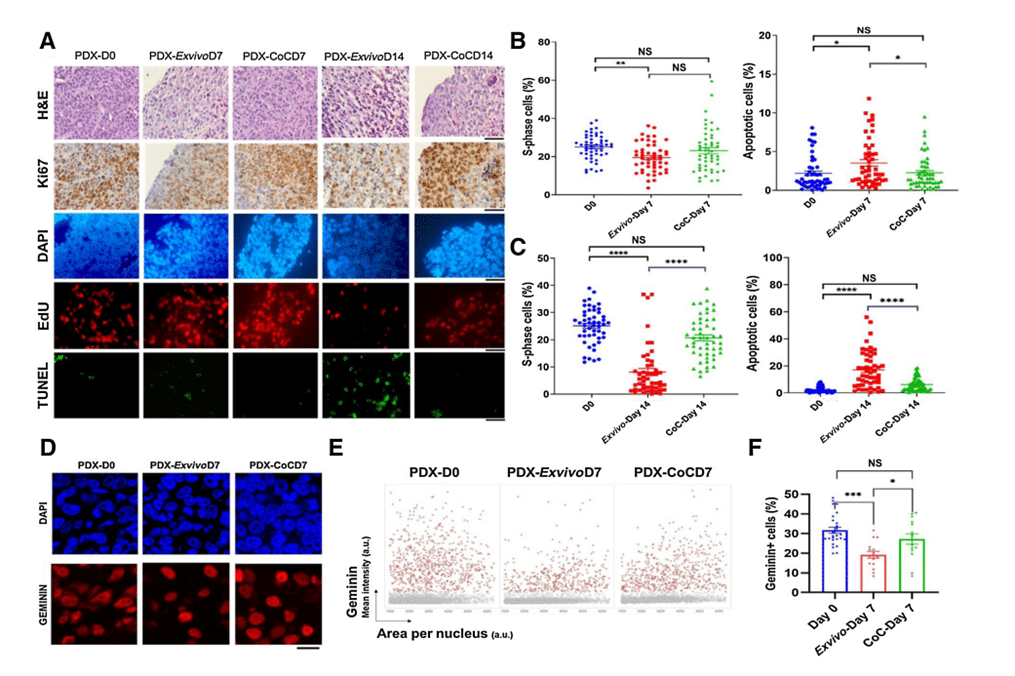
Figure 4: Breast PDX tumor tissue slices cultured in ex vivo condition and in CoC device for up to 14 days. A, Quantification of the fraction of EdU-positive and TUNEL-positive cells for 5 breast PDX tissue slices cultured for up to day 7 (B) and day 14 (C). D, Representative image showing breast PDX tumors labeled with geminin (red nuclei) and DAPI (blue nuclei). E, QIBC analysis of three independent breast PDX tumors with more than 3,000 cells analyzed for each are shown in each condition. F, Quantification of geminin-positive cells showed CoC at day 7 had similar cell proliferation profile as in day 0 than ex vivo condition.
Gene expression analysis
To assess the impact of CoC culture on gene expression changes, analysis using RT-PCR and whole transcriptome sequencing was performed. Tumor-specific gene pathways in PDX breast tumors cultured ex vivo and in the Cancer-on-a-Chip platform were examined. Surprisingly, there were no statistically significant changes in these pathways, suggesting minimal alterations in tumor growth characteristics under both conditions. Next, whole transcriptome sequencing of PDX breast tumors was performed. Genes were identified as differentially expressed on day 7 ex vivo and on days 7 and 14 under CoC culture conditions. 150 human genes are differentially expressed in day 7 ex vivo tumor slices, far more than the 30 human genes differentially expressed in day 7 CoC and the 14 human genes in day 14 CoC.
To understand the reasons for the differences observed, various tests studying cell cycle progression and apoptosis were carried out. They led to the conclusion that ex vivo culture conditions induced greater immune activation and DNA damage after 7 days, making the CoC system a more accurate representation of the original tumor and the preferred choice for studying responses to therapies.
Conclusion
The researchers developed a microfluidic CoC platform capable of maintaining cell viability, proliferation and tissue structure in breast cancer PDX slices for at least 14 days. This platform successfully predicted responses to cisplatin therapy for breast cancer and antiandrogen therapy for PDX prostate cancer tumor slices. To fully establish its potential as an in vitro diagnostic test for therapy selection, it will require clinical validation using biopsies from patients receiving the same chemotherapy.Although the current study has focused on PDX models of breast and prostate cancer, this Cancer-on-a-Chip platform also holds promise for other solid tumors. Its ease of use and small footprint make it a versatile tool for ex vivo studies, including functional genomics, drug screening and personalized medicine research.
Expertises & Resources
-
Expert Reviews: Basics of Microfluidics Microfluidic pressure control for organ-on-a-chip applications: A comprehensive guide Read more
-
Microfluidic Application Notes Long-term fluid recirculation system for Organ-on-a-Chip applications Read more
-
Microfluidics case studies CNRS/UTC: study of a liver-on-a-chip model Read more
-
Interviews & Testimonials Panel Discussion & Interviews – Microfluidics & Organ-On-Chips Read more
-
Microfluidics White Papers Microfluidic white paper – A review of Organ on Chip Technology Read more
-
Microfluidic Application Notes Peristaltic Pump vs Pressure-Based Microfluidic Flow Control for Organ on Chip applications Read more
Related Webinars
Related products
References
[1] Microfluidic white paper – A guide to Organs-on-Chips technology, Fluigent
[2] Naipal KA, Verkaik NS, Sanchez H, van Deurzen CH, den Bakker MA, Hoeijmakers JH, et al. Tumor slice culture system to assess drug response of primary breast cancer. BMC Cancer 2016;16:78
[3] van Weerden WM, van Steenbrugge GJ, van Kreuningen A, Moerings EP, de Jong FH, Schr€oder FH. Assessment of the critical level of androgen for growth response of transplantable human prostatic carcinoma (PC-82) in nude mice. J Urol 1991;145:631–4