Microfluidics for vaccine development
The WHO estimated worldwide demand for vaccines at 3.5 billion doses in 2018, up approximately 25% compared to 2017.1 Alongside the demand for existing vaccines, the appearance of new viruses such as Ebola and COVID-19 has led to even more critical levels of demand. A report from IPBES also states that increasing exploitation of the environment will result in more frequent occurrence of new pandemics.2
Together, all of these factors contribute to increased pressure on vaccine development time and cost. Offering control over particle size, molecular concentrations, microenvironment, compartmentalization and high-throughput screening, microfluidics for vaccine development is growing fast and drawing increasing interest in the scientific community.
Fluigent’s mission is to develop solutions and technologies to support researchers and industry experts conducting microfluidic experiments under excellent conditions.
Technological advances for vaccine development
First-generation vaccines are based on whole organisms, in either live attenuated or killed forms. Second-generation vaccines consist of protein components derived from the organism. Today, third-generation vaccines are being developed. These nucleic acid vaccines are based on the genetic material of the infectious organism. DNA vaccines are examples of third generation vaccines. Clinical trials for DNA vaccines to prevent HIV are currently underway.
During the COVID-19 pandemic, the first vaccines to reach clinical trials were based on viral vector and nucleic acid technologies. One of the most promising vaccine candidates was based on mRNA, as it offers a cost-effective, easy to manufacture, adaptable solution and induces both a cellular and a humoral response.
Scientific advances are becoming more and more critical in the success of fundamental vaccine research, vaccine development, and diagnostics. New methods such as microfluidics for vaccine research can be used to improve development processes.
Microfluidics in vaccine research and development
Efficient delivery of mRNA and DNA into target cells in vivo is a major challenge. Today, many vaccines are encapsulated into nanoparticles, specifically lipid nanoparticles (LNP) such as liposomes, for improved delivery efficiency. The first COVID-19 vaccines to reach the market are mRNA vaccines encapsulated in LNP.
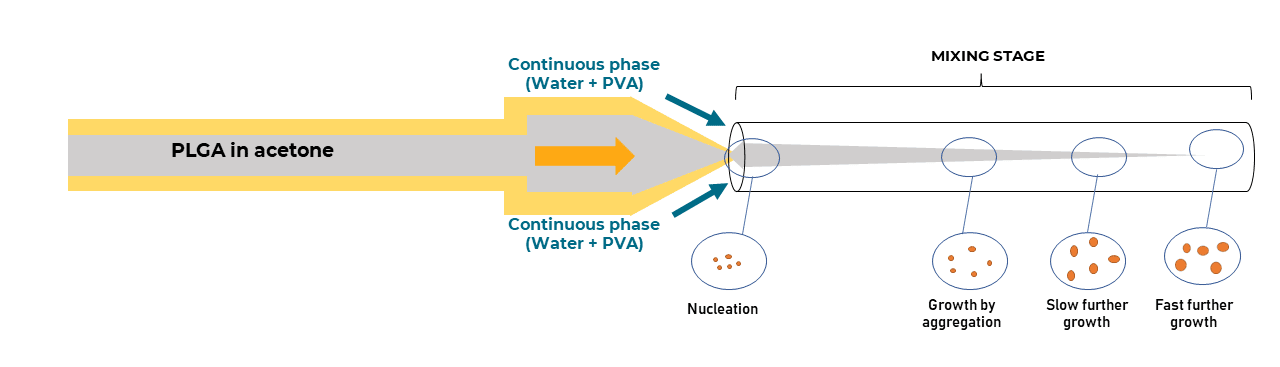
Size and size distribution are essential properties in determining the clinical success of nanocarriers. Recently, improvements have been made in the development of microfluidic production methods, in which LNP formation occurs within a confined microenvironment with spontaneous self-assembly of the phospholipids as a sphere to minimize surface energy. These methods have demonstrated higher control over the physical properties of the end product, particularly in terms of liposome size and size distribution.
Examples of microfluidics applications in vaccine development
Vaccine adjuvant production
Among the various applications of microfluidics in vaccine development, adjuvant production is on the front line. Adjuvants are used to amplify the recipient’s specific immune responses against pathogen infection. A new generation of adjuvants is being developed to meet the demands of improved antigen-specific responses with less toxicity. For example, Konishi et al. reported the first synthesis of a series of saponins, a vaccine adjuvant, using a microfluidic mixer.3

High throughput screening
Flow cytometry for single-virus analysis shows high potential in virology, vaccine development, and antiviral research. However, such analysis can be difficult to perform because fluorescence-activated cell sorting (FACS), the standard technique in modern biology, does not allow for single-virus measurements due to viruses’ small size.

The Fluigent double emulsion production station is a robust and complete system for producing outstanding monodispersed double emulsions with a single device.
Droplet-based microfluidics combined with next-generation sequencing can allow for high-throughput screening of virus particles. For instance, Chaipan et al. have developed a droplet-based microfluidic platform to screen single virus particles for optimal antigenic features of vaccine candidates and demonstrated the platform’s use by screening HIV particles.4
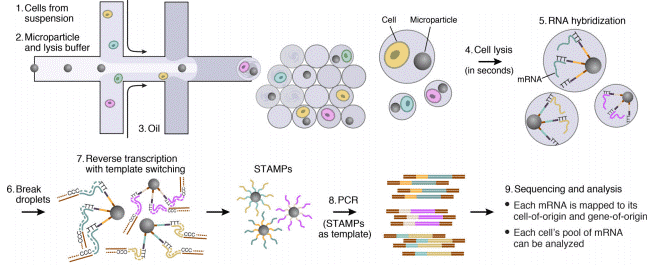
The Drop-Seq protocol is a high-throughput method that enables sequencing of mRNA from a large number of cells.
Virus detection and analysis
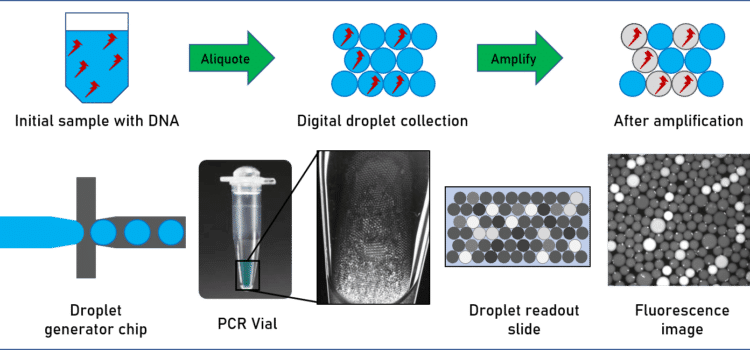
Finally, virus detection applications are of particular interest in vaccine development. The quantity of virus in a sample is typically quite low, and diluted in a mix of all kinds of proteins, DNA and RNA. An amplification step is therefore necessary to increase the quantity of detectable RNA. Polymerase chain reaction (PCR), and other PCR-related methods such as quantitative PCR (qPCR), amplify and quantify DNA and RNA for subsequent analysis. In contrast to standard PCR, digital PCR (dPCR) divides the sample into subsamples in the pico- to nanoliter range, allowing for more reliable collection and more sensitive measurement of nucleic acid quantities.
For example, Ahrberg et al. have developed an easy-to-use microfluidic dPCR device to amplify and quantify complementary deoxyribonucleic acid (cDNA) samples for the H7N9 influenza virus.5 In another study, researchers from the California Institute of Technology and MIT developed a microfluidic dPCR approach to physically link single bacterial cells harvested from a natural environment with a viral marker gene for examining virus-bacterium interactions in many different environments.6
In-vitro cell analysis in vaccine research
Cell culture is at the center of many biological experiments, as it is the first step in most in-vitro analyses. The conditions in which the cells are cultivated determine the culture’s growth potential, protein secretion, and cell-cell signaling, and bulk cultures sometimes fail to effectively control these parameters.
Microfluidics provide unique capabilities to control the cellular microenvironment and present cells with mechanical and biochemical signals in a more physiologically relevant context. Microfluidic chips offer an ideal microenvironment to study the molecular- and cellular-scale activities that underlie human organ function and identify new therapeutic targets in vitro. For instance, Markus et al. used microfluidic cell culture chambers to analyze the hosting of varicella-zoster virus in human embryonic stem cell-derived neurons.7
References
- World Health Oganization. Global vaccine market report. 14 p (2019).
- Luthando Dziba, Isabel Sousa Pinto, Judith Fisher, K. T. IPBES Workshop on biodiversity and pandemics. (2020).
- Konishi, N. et al. Synthesis of Bisdesmosidic Oleanolic Acid Saponins via a Glycosylation-Deprotection Sequence under Continuous Microfluidic/Batch Conditions. J. Org. Chem. 82, 6703–6719 (2017).
- Chaipan, C. et al. Single-Virus Droplet Microfluidics for High-Throughput Screening of Neutralizing Epitopes on HIV Particles. Cell Chem. Biol. 24, 751-757.e3 (2017).
- Ahrberg, C. D., Lee, J. M. & Chung, B. G. Microwell Array-based Digital PCR for Influenza Virus Detection. Biochip J. 13, 269–276 (2019).
- Arbel D. Tadmor, Elizabeth A. Ottesen, Jared R. Leadbetter, and R. P. Probing Individual Environmental Bacteria for Viruses by Using Microfluidic Digital PCR. Science (80-. ). 23, 1–7 (2012).
- Markus, A., Lebenthal-Loinger, I., Yang, I. H., Kinchington, P. R. & Goldstein, R. S. An In Vitro Model of Latency and Reactivation of Varicella Zoster Virus in Human Stem Cell-Derived Neurons. PLoS Pathog. 11, 1–22 (2015).