High encapsulation efficiency
Drop-Seq Pack
[ODROPSEQFPCK]Complete system for starting Drop-Seq experiments
Droplet sequencing is a low-cost, high-throughput platform to profile thousands of cells by encapsulating them into individual droplets. Uniquely barcoded mRNA capture microparticles and cells are coconfined through a microfluidic device within the droplets where they undergo cell lysis and RNA hybridization.
With our droplet-sequencing pack, you will be able to perform optimized Drop-Seq experiments in Next-Generation Sequencing (NGS). The Fluigent Drop-Seq pack allows better reproducibility and control of both single cells and beads encapsulation.
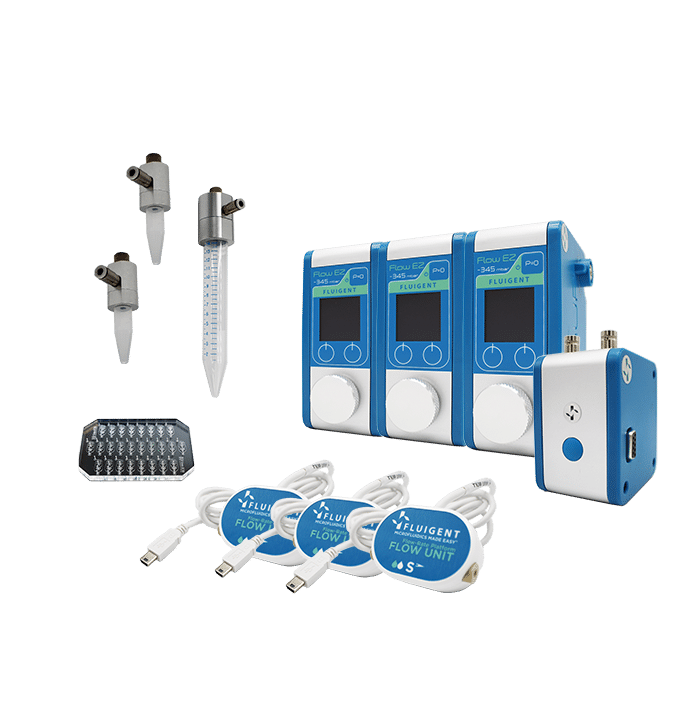

- Efficient
- Gain time
Faster than standard Drop-Seq protocol
- Reproducible
High monodispersity (2%)
Features of our drop-seq pack
Latest design
Each droplet generation device, including the droplet-sequencing pack is based on the design recommended in the latest McCarroll lab Drop-Seq protocol, ensuring the best chances of success.
Fluigent devices
Precision engineered robust devices durable over a wide range of pressures, temperatures, and flow rates Last
23 designs per chip
Provides value for money in a chip that lasts. When the life of one device is depleted, simply move onto the next one
Produce monodisperse droplets
Reliable and consistent generation of droplets of optimal size for Droplet sequencing
Efficient production of transcript libraries
Superior design promotes optimal mixing of component fluids, thereby minimizing bead shearing or premature lysis of cells and mRNA release
Why use pressure for Droplet sequencing?
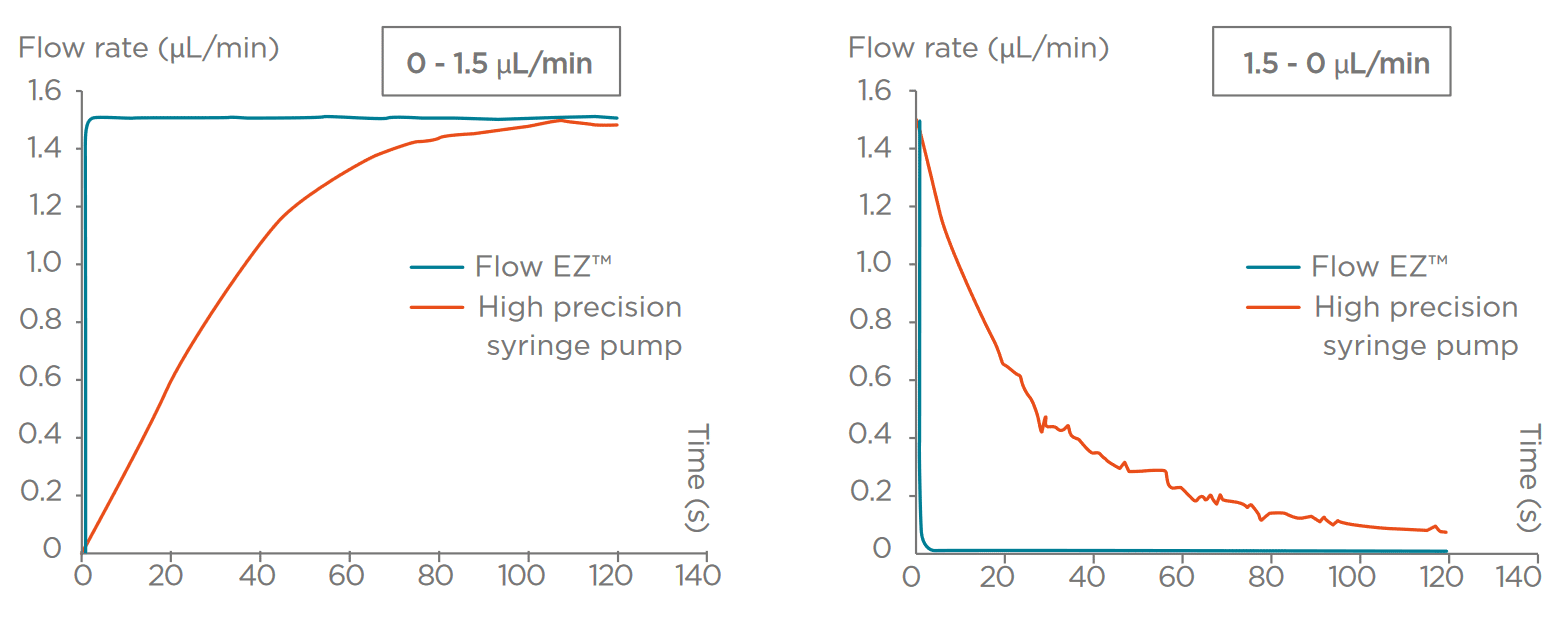
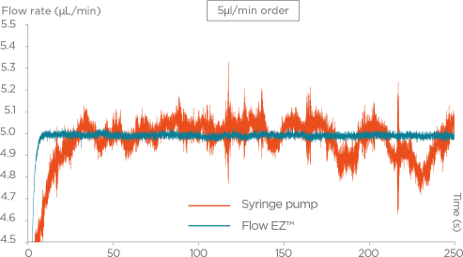
While the original protocol was developed using syringe pumps, in the field of droplet sequencing, pressure-controlled systems have a few advantages.
First, in terms of raw performance, pressure-driven systems are faster to set up, easier to control, and more stable over time, which allows for an emulsion of better quality: the droplet size will be more homogeneous, and start/stop populations will be smaller. This leads to better segregation of the cells, and less reagent use and less sample loss. Therefore, this allows for more efficient and optimal droplet sequencing.
As the sample and beads need to be agitated throughout the experiment, it is usually done with a bulky stirring bar for syringe pumps. This leads to a dead volume that will not be injected into the chip. Pressure-driven setups can be run using more conventional containers that make it possible to use an external agitation system, such as a standard lab vortexing system.
Using Fluigent Drop-Seq pack experiment with pressure controller allow then to:
- Gain time: (less than 1 minute to obtain droplet compared to Macosko’s protocol, few minutes).
- Avoid losing reagents (cells or beads) during transition time.
- Have better control and avoid the problems that could appear with a syringe pump. For instance, using the droplet-sequencing pack prevents the backflow of the beads inside the cell’s channel which could drastically modify the experiment.
Main products of the package
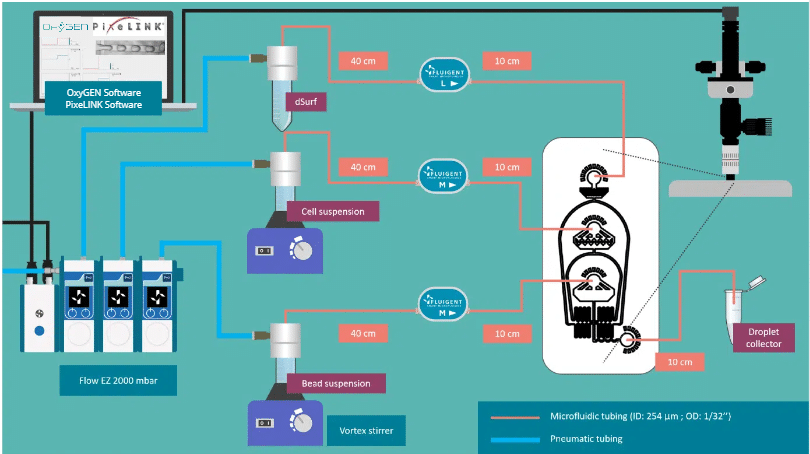
Specifications
| LineUp Flow EZ pressure controller (2 bar) x3 |
| FLOW UNIT M x2 |
| FLOW UNIT L x1 |
| LineUp LINK Module (software control) x1 |
| P-CAP 15 mL x1 |
| P-CAP 2 mL x2 |
| Drop-Seq chip |
| LineUp Supply kit x1 |
| Tubing & fitting kit x1 |
OxyGEN
| Control in real-time, protocol automation, data record and export |
| ver. 2.2.0.0 or more recent |
Software Development Kit
| Custom software application |
| ver. 22.2.0.0 or more recent |
Expertise & resources
Related products
Accessories
Kits
References
[1] Théry M, Bornens M, Get round and stiff. 2008, HFSP J, 2(2):65-71.
[2] Tavares S et al, actin stress fiber organization promotes cell stiffening and proliferation of pre-invasive breast cancer cells. 2017, Nat Commun. 8:15237.
[3] Guo Q et al, Microfluidic biomechanical assay for red blood cells parasitized by Plasmodium falciparum. 2012, Lab Chip; 12(6):1143-50.
[4] Yanez LZ et al, human oocyte developmental potential is predicted by mechanical properties within hours after fertilization, 2016, Nat Commun. 7:10809